\n
Jump to:
\n
\n
\n
\n Bio
\n
“The Lands
cape of the Heritable Cancer Genome”\n

\n
Giovanni Stracquadanio is an UKRI EPSRC
fellow\, Senior Lecturer (Associate Professor) in Synthetic Biology and c
o-director of the Edinburgh Genome Foundry (EGF). He group is interested i
n understanding the molecular mechanisms underpinning complex phenotypes a
nd diseases using two of the most disruptive technologies available: synth
etic biology and machine learning. His long-term goal is to reverse-engine
er biological systems to develop generative algorithms to design\, build a
nd test biological agents for addressing healthcare problems\, such as rar
e metabolic diseases and cancer\, and industrial biotechnology challenges\
, like de-novo enzyme engineering.
\n
Dr Stracquadanio obtained a PhD
in Informatics from the University of Catania (Italy) in 2010\, working o
n global optimisation methods for protein structure prediction and metabol
ic engineering. He then received postdoctoral training in synthetic biolog
y in Joel Bader and Jef Boeke labs at the Johns Hopkins University working
on the synthetic yeast genome.
\n
Dr Stracquadanio was a main contri
butor to the Synthetic Yeast (Sc2.0) genome project\, pioneering algorithm
s and developing software at the foundation of the first synthetic eukaryo
tic genome. He has also developed tools used in large-scale synthesis proj
ects\, streamlining chromosomes engineering and the assembly of biological
pathways.
\n
In 2014\, he moved to the Ludwig Institute for Cancer R
esearch at the University of Oxford UK to work on cancer genetics\, in Gar
eth Bond’s lab\; here\, he focused on studying how high-frequency inherite
d p53 mutations affect the risk of cancer and response to treatment using
statistical genetics methods and cell line models.
\n
In 2016\, prior
to joining the University of Edinburgh\, Dr Stracquadanio was an assistan
t professor at the School of Computer Science and Electronic Engineering o
f the University of Essex\, where he received the Wellcome Trust Seed Awar
d in Science to study metabolic switching in renal cell carcinoma.
\n
In 2021\, he was awarded the UKRI EPSRC fellowship to design and manufact
ure enzyme replacement therapies for Fabry’s disease using generative deep
learning and yeast. He also collaborates with industrial stakeholders in
the biotechnology field\, such as Fujifilm Diosynth\, on protein design\,
codon optimisation and lab automation.
\n
\n
Dr Stracquadanio
has authored more than 40 research articles published in international pee
r-reviewed journals\, including Science\, Nature Rev. Cancer\, Cancer Rese
arch and PNAS. He also serves as Associate Editor for BMC Genomics and as
reviewer and panel member for EPSRC\, BBSRC\, MRC and FLF. Since 2021\, he
is also a member of the EPSRC Peer Review Associate College.
\n
.
\n
\n
To join the live event please request the link by emailing
: icm@jhu.edu
\n
<
/p>\n
 Recording
Recording
\n
\n
\n Ab
stract
\n
\n
“The Landscape of the Heritable
Cancer Genome”
\n
\n
\n
\n
\n
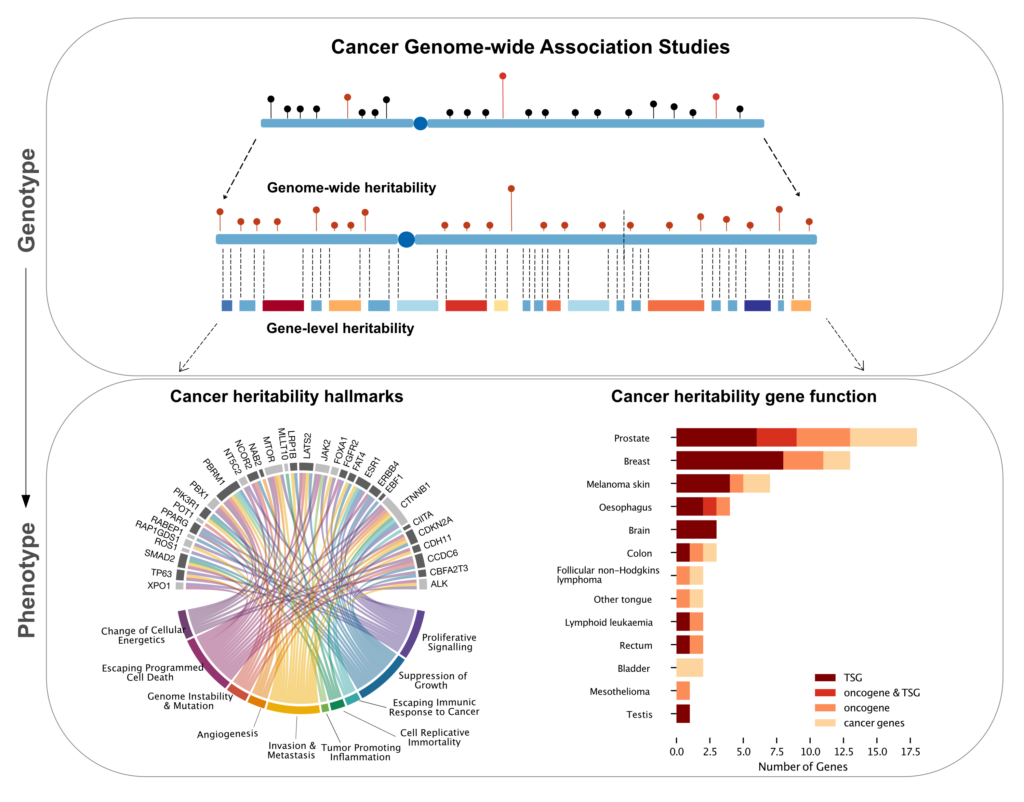
Genome-wide asso
ciation studies (GWAS) have found hundreds of single nucleotide polymorphi
sms (SNPs) associated with increased risk of cancer. However\, the amount
of heritable risk explained by SNPs is limited\, leaving most of cancer he
ritability unexplained. Tumor sequencing projects have shown that causal m
utations are enriched in genic regions. We hypothesized that SNPs located
in protein coding genes and nearby regulatory regions could explain a sign
ificant proportion of the heritable risk of cancer.
\n
To perform gen
e-level heritability analysis\, we developed a new method\, called Bayesia
n Gene HERitability Analysis (BAGHERA)\, to estimate the heritability expl
ained by all genotyped SNPs and by those located in genic regions using GW
AS summary statistics [1]. BAGHERA-based analysis of 38 cancers reported i
n the UK Biobank showed that SNPs explain at least 10% of the heritable ri
sk for 14 of them\, including late onset malignancies. We then identified
1\,146 genes\, called cancer heritability genes (CHGs)\, explaining a sign
ificant proportion of cancer heritability. CHGs were involved in hallmark
processes controlling the transformation from normal to cancerous cells. I
mportantly\, 60 of them also harbored somatic driver mutations\, and 27 ar
e tumor suppressors. Evidence for a causal role of CHGs was shown in testi
cular cancer\, where a cluster of SNPs in the KITLG CHG is responsible for
p53-mediated responses to genotoxic therapies [2].
\n
Our results su
ggest that germline and somatic mutation information could be exploited to
identify subgroups of individuals at higher risk of cancer in the broader
population and could prove useful to establish strategies for early detec
tion and cancer surveillance.
\n
 Recording
Recording
\n<
/div>\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4983373@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series
CONTACT:Mishka Colombo\; 4105164116\; mishka@jhu.edu\; https://icm.jhu.edu/
seminar-series/
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Mechanistic and Data-Driven Dissection of Cell Communication Thr
ough Tensor Methods”\n\nDr. Aaron Meyer is an Assistant Professor of Bioen
gineering and Bioinformatics at the University of California\, Los Angeles
. He previously received his B.S. in Bioengineering from UCLA\, his Ph.D.
in Biological Engineering from the Massachusetts Institute of Technology\,
and then was an independent fellow at the Koch Institute for Integrative
Cancer Research. Dr. Meyer’s research aims to develop new computational me
thods\, deeply integrated with experiments\, to understand intercellular c
ommunication and how it can be therapeutically manipulated. He primarily f
ocuses on combinations of mechanistic and data-driven modeling as a soluti
on to our incomplete knowledge of cellular pathways\, and applications in
cancer and immunity where models can have immediate therapeutic impact. Hi
s work has been recognized by the NIH Director’s Early Independence Award\
, a Siebel Scholars award\, a Hellman Fellowship\, the UCLA Faculty Career
Development Award\, the Northrop Grumman Excellence in Teaching Award\, a
nd named as Ten to Watch by the Amgen Foundation.\n \nTo join the live eve
nt please request the link by emailing: icm@jhu.edu\n \nRecording\n\n
\n Abstract \n\n“Mechanistic and Data-Driven Dissection of Cell
Communication Through Tensor Methods”\n \n\n \n \nStudies of even simple
cell responses to their environment are hindered by how responses are mult
i-dimensional. For example\, a simple receptor-ligand pathway can display
differing responses based on timescale\, cell type\, stimulation\, type of
response measured\, and context. Interrogating and manipulating these sys
tems is thus almost always constrained by an incomplete view of the overal
l pathway.\n \nLike how principal component analysis uses a low-rank appro
ximation for dimensionality reduction of matrix-structured data\, tensor g
eneralizations provide solutions for pattern recognition in data with a hi
gher-dimensional structure. Using several recent and unpublished applicati
ons\, including engineering cell-type selective IL-2 therapies\, serology
analysis\, and clinical multi-omic studies of infection response\, I will
describe some of the unique benefits of tensor-based analysis and the biol
ogical discoveries it has revealed. Specifically\, tensor approximations e
nable more effective dimensionality reduction\, separation of dimension-sp
ecific effects\, and a natural\, flexible solution to data integration. Fi
nally\, I will discuss some of the reasons tensor-based methods remain lim
ited in their application to molecular biology. Resolving these limitation
s\, and applying tensor methods in a more widespread manner\, will help pr
ovide a complete view of cellular communication.\nRecording
DTSTART;TZID=America/New_York:20220201T103000
DTEND;TZID=America/New_York:20220201T113000
LOCATION:Zoom
SEQUENCE:0
SUMMARY:Mechanistic and Data-Driven Dissection of Cell Communication Throug
h Tensor Methods
URL:https://icm.jhu.edu/events/mechanistic-and-data-driven-dissection-of-ce
ll-communication-through-tensor-methods/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2022/01/a
aron-meyer-headshot-1-1024x1024.jpg\;111\;111\,medium\;https://icm.jhu.edu
/wp-content/uploads/2022/01/aaron-meyer-headshot-1-1024x1024.jpg\;111\;111
\,large\;https://icm.jhu.edu/wp-content/uploads/2022/01/aaron-meyer-headsh
ot-1-1024x1024.jpg\;111\;111\,full\;https://icm.jhu.edu/wp-content/uploads
/2022/01/aaron-meyer-headshot-1-1024x1024.jpg\;111\;111
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Mechanist
ic and Data-Driven Dissection of Cell Communication Through Tensor Methods
”
\n

\n
Dr. Aaron Meyer is an Assist
ant Professor of Bioengineering and Bioinformatics at the University of Ca
lifornia\, Los Angeles. He previously received his B.S. in Bioengineering
from UCLA\, his Ph.D. in Biological Engineering from the Massachusetts Ins
titute of Technology\, and then was an independent fellow at the Koch Inst
itute for Integrative Cancer Research. Dr. Meyer’s research aims to develo
p new computational methods\, deeply integrated with experiments\, to unde
rstand intercellular communication and how it can be therapeutically manip
ulated. He primarily focuses on combinations of mechanistic and data-drive
n modeling as a solution to our incomplete knowledge of cellular pathways\
, and applications in cancer and immunity where models can have immediate
therapeutic impact. His work has been recognized by the NIH Director’s Ear
ly Independence Award\, a Siebel Scholars award\, a Hellman Fellowship\, t
he UCLA Faculty Career Development Award\, the Northrop Grumman Excellence
in Teaching Award\, and named as Ten to Watch by the Amgen Foundation.
\n
\n
To join the live event please request the link by emailing
: icm@jhu.edu
\n
<
/p>\n
 Recording
Recording
\n
\n
\n Abstract
\n
\n
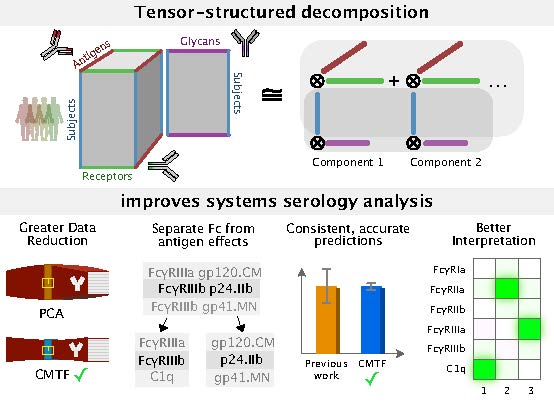
“Mechanistic and Data-Driven Dissection o
f Cell Communication Through Tensor Methods”<
/span>
\n
\n
\n
\n
\n
Studies of even simple cell responses to their envi
ronment are hindered by how responses are multi-dimensional. For example\,
a simple receptor-ligand pathway can display differing responses based on
timescale\, cell type\, stimulation\, type of response measured\, and con
text. Interrogating and manipulating these systems is thus almost always c
onstrained by an incomplete view of the overall pathway.
\n
\n
Like how principal component analysis uses a low-rank approximation for d
imensionality reduction of matrix-structured data\, tensor generalizations
provide solutions for pattern recognition in data with a higher-dimension
al structure. Using several recent and unpublished applications\, includin
g engineering cell-type selective IL-2 therapies\, serology analysis\, and
clinical multi-omic studies of infection response\, I will describe some
of the unique benefits of tensor-based analysis and the biological discove
ries it has revealed. Specifically\, tensor approximations enable more eff
ective dimensionality reduction\, separation of dimension-specific effects
\, and a natural\, flexible solution to data integration. Finally\, I will
discuss some of the reasons tensor-based methods remain limited in their
application to molecular biology. Resolving these limitations\, and applyi
ng tensor methods in a more widespread manner\, will help provide a comple
te view of cellular communication.
\n
Recording
\n
\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4983944@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series
CONTACT:Mishka Colombo\; 4105164116\; mishka@jhu.edu\; https://wse.zoom.us/
rec/play/DvxOSxUu_9WVBbRq34jXYwNn3JQT-jg0CLF9MWN4cZ4c7AZ2taBpdKQq4ER9C1RSv
13DEpkjE9liT1qH.ebm19KbcRLY59FZG?continueMode=true&_x_zm_rtaid=VygAu5OITom
e1kFCWhsfmw.1648580481104.a2f113c9ae4cb6de3e9f7f07dc46450b&_x_zm_rhtaid=42
3
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Mechanistic Modeling of Signal Transduction and Dynamic Cell Mor
phologies”\n\nDr. Meier-Schellersheim’s research group has been developing
tools for quantitative computational image analysis and mechanistic model
ing of cellular behavior for several years now. They were able to develop
computational methods and tools that permit the simulation of cellular sig
naling networks embedded into dynamic\, realistic 3D morphologies to take
into account the coupling between cellular biochemistry and morphological
dynamics. The simulation platform Simmune was the first modeling software
to permit this degree of realism and we have continuously been improving i
ts capabilities with regard to the size of multi-cellular systems that can
be simulated and parameter scans that can be performed on distributed com
puter systems. Simmune can model cells that express receptors for chemosen
sing and adhesion on their surface that react to receptor mediated stimuli
by adjusting their geometry to adhere to extracellular structures or by d
irected migration in response to chemotactic signals. Although the couplin
g between biochemistry and cell motion based on Potts model rules is pheno
menological applying such simulation techniques to explore the role cell-c
ell and cell-matrix interactions may play in complex 3D structures is a fi
rst step towards understanding how chemical and mechanical cues regulate c
ellular migration. Dr. Meier-Schellersheim’s group develops and applies qu
antitative image analysis tools to extract the information needed for deta
iled spatially resolved simulations directly in an unbiased and automated
manner from image data. Having been part of the Laboratory of Systems Biol
ogy for several years now\, his group acquired considerable experience in
coordinating interdisciplinary work and in handling large heterogeneous da
ta sets.\nIn addition to exploring the interplay between cell morphology a
nd cellular responses towards stimuli\, Dr. Meier-Schellersheim’s group de
velops and applies tools that can perform systematic analyses of the behav
ior of computational models of cellular signaling pathways. These tools ca
n identify which elements of a pathway model are responsible for reproduci
ng specific features in the experimentally observed cellular behavior.\n
\nTo join the live event please request the link by emailing: icm@jhu.edu
\n \n▶RECORDING\n\n \n Abstract \n\n“Mechanistic Modeling o
f Signal Transduction and Dynamic Cell Morphologies”\n \n\n \n \nDetailed
mechanistic models of cellular signaling pathways have the advantage that
the conclusions they allow us to draw can be tested with little ambiguity.
However\, building such models typically requires more data and involves
far more parameters than phenomenological approaches do. Using examples fr
om cytokine and growth factor signaling\, I will discuss some recent progr
ess in applying detailed modeling tools and describe what can be learned f
rom models whose parameters cannot be uniquely determined. Then\, I will s
how how detailed models of intracellular biochemistry can be linked to a r
ealistic treatment of morphological dynamics using a novel approach for re
presenting cellular surfaces.\n \n \n▶RECORDING
DTSTART;TZID=America/New_York:20220329T103000
DTEND;TZID=America/New_York:20220329T113000
LOCATION:Zoom @ email mishka@jhu.edu for link
SEQUENCE:0
SUMMARY:Mechanistic Modeling of Signal Transduction and Dynamic Cell Morpho
logies
URL:https://icm.jhu.edu/events/mechanistic-modeling-of-signal-transduction-
and-dynamic-cell-morphologies/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2022/03/M
artin-Meier-Schellersheim.jpg\;107\;132\,medium\;https://icm.jhu.edu/wp-co
ntent/uploads/2022/03/Martin-Meier-Schellersheim.jpg\;107\;132\,large\;htt
ps://icm.jhu.edu/wp-content/uploads/2022/03/Martin-Meier-Schellersheim.jpg
\;107\;132\,full\;https://icm.jhu.edu/wp-content/uploads/2022/03/Martin-Me
ier-Schellersheim.jpg\;107\;132
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Mechan
istic Modeling of Signal Transduction and Dynamic Cell Morphologies”
\n

\n
Dr. Meier-Schellersheim’s research gr
oup has been developing tools for quantitative computational image analysi
s and mechanistic modeling of cellular behavior for several years now. The
y were able to develop computational methods and tools that permit the sim
ulation of cellular signaling networks embedded into dynamic\, realistic 3
D morphologies to take into account the coupling between cellular biochemi
stry and morphological dynamics. The simulation platform Simmune was the f
irst modeling software to permit this degree of realism and we have contin
uously been improving its capabilities with regard to the size of multi-ce
llular systems that can be simulated and parameter scans that can be perfo
rmed on distributed computer systems. Simmune can model cells that express
receptors for chemosensing and adhesion on their surface that react to re
ceptor mediated stimuli by adjusting their geometry to adhere to extracell
ular structures or by directed migration in response to chemotactic signal
s. Although the coupling between biochemistry and cell motion based on Pot
ts model rules is phenomenological applying such simulation techniques to
explore the role cell-cell and cell-matrix interactions may play in comple
x 3D structures is a first step towards understanding how chemical and mec
hanical cues regulate cellular migration. Dr. Meier-Schellersheim’s group
develops and applies quantitative image analysis tools to extract the info
rmation needed for detailed spatially resolved simulations directly in an
unbiased and automated manner from image data. Having been part of the Lab
oratory of Systems Biology for several years now\, his group acquired cons
iderable experience in coordinating interdisciplinary work and in handling
large heterogeneous data sets.
\n
In addition to exploring the inter
play between cell morphology and cellular responses towards stimuli\, Dr.
Meier-Schellersheim’s group develops and applies tools that can perform sy
stematic analyses of the behavior of computational models of cellular sign
aling pathways. These tools can identify which elements of a pathway model
are responsible for reproducing specific features in the experimentally o
bserved cellular behavior.
\n
\n
To join the live event please
request the link by emailing: icm@jh
u.edu
\n
\n
▶RECORDING
\n
\n
\n Abstract
\n
\n
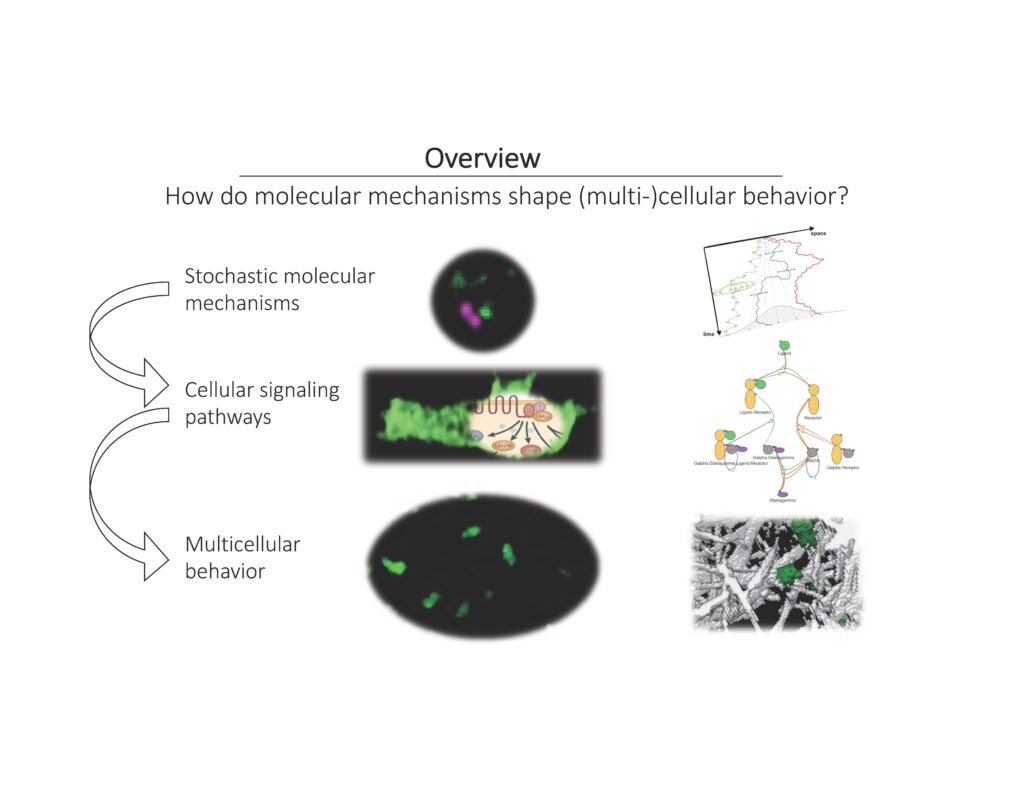
“Mechanistic Modeling of Signal Transduction and Dynamic Cell Morphol
ogies”
\n
\n
\n
\n
\n
Detailed mechanistic models of cellular signaling pathways
have the advantage that the conclusions they allow us to draw can be teste
d with little ambiguity. However\, building such models typically requires
more data and involves far more parameters than phenomenological approach
es do. Using examples from cytokine and growth factor signaling\, I will d
iscuss some recent progress in applying detailed modeling tools and descri
be what can be learned from models whose parameters cannot be uniquely det
ermined. Then\, I will show how detailed models of intracellular biochemis
try can be linked to a realistic treatment of morphological dynamics using
a novel approach for representing cellular surfaces.
\n
\n
<
/p>\n
▶RECORDING
\n
\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984318@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series\,Events
CONTACT:Mishka Colombo\; 4105164116\; mcolomb4@jhu.edu\; email mcolomb4@jhu
.edu for link
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Towards a New Comprehensive Human Gene Catalogue”\n \nMihaela Pe
rtea is an Associate Professor in the Department of Biomedical Engineering
at Johns Hopkins University. She received her B.S. and M.S. degrees in Co
mputer Science from University of Bucharest in Romania\, and her Ph.D in C
omputer Science from the Johns Hopkins University School of Engineering. D
r. Pertea’s work in computational biology draws upon techniques and data f
rom multiple disciplines\, including computer science and molecular biolog
y\, genetics\, biotechnology\, and statistics. Her work has focused on com
putational gene finding and sequence pattern recognition and she has devel
oped several open-source gene finders that were used for the annotation of
the genomes of Plasmodium falciparum (malaria parasite)\, Arabidopsis tha
liana\, rice\, Aspergillus fumigatus\, Cryptococcus neoformans\, and other
s. A major focus of her current research is on developing innovative and e
fficient methods to analyze large DNA and RNA sequence data in order to pr
ovide a genome-scale understanding of cellular function. Dr. Pertea believ
es that the principled use of algorithms from other fields\, adapted to th
e problems of computational biology and coupled with careful software engi
neering and high performance computing\, has the potential to make a signi
ficant impact in the life sciences. She has published over 50 scientific p
apers that have received more than 30\,000 citations to date.\n \nTo join
the live event please request the link by emailing: icm@jhu.edu\n \n▶RECOR
DING \n\n \n Abstract \n \n“Towards a New Comprehensive Hum
an Gene Catalogue”\n\n \n\n \n \nA huge and still-growing number of geneti
c studies depend on the human gene catalogue\, including thousands of expe
riments each year and an enormous investment of time and effort. However\,
despite its critical role in biomedical research\, the human gene list is
still incomplete and\, in many ways\, unstable. The widespread use of RNA
sequencing technology over the past decade has allowed scientists to disc
over a far larger and richer repertoire of genes and transcripts than prev
iously known. Our own recent efforts led to the creation of a new human g
ene catalogue\, called CHESS\, that we built using a very large collection
of nearly 10\,000 RNA-seq experiments from 31 tissues\, all sequenced as
part of the GTEx project. Processing this large amount of data was one of
the most challenging tasks\, made possible by the computational efficiency
of StringTie\, a transcriptome assembler we developed in our lab.\n \n▶RE
CORDING
DTSTART;TZID=America/New_York:20220426T103000
DTEND;TZID=America/New_York:20220426T113000
LOCATION:Zoom
SEQUENCE:0
SUMMARY:Towards a New Comprehensive Human Gene Catalogue
URL:https://icm.jhu.edu/events/towards-a-new-comprehensive-human-gene-catal
ogue/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2022/03/M
ichaela-Pertea-1.jpg\;137\;137\,medium\;https://icm.jhu.edu/wp-content/upl
oads/2022/03/Michaela-Pertea-1.jpg\;137\;137\,large\;https://icm.jhu.edu/w
p-content/uploads/2022/03/Michaela-Pertea-1.jpg\;137\;137\,full\;https://i
cm.jhu.edu/wp-content/uploads/2022/03/Michaela-Pertea-1.jpg\;137\;137
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Toward
s a New Comprehensive Human Gene Catalogue”
\n

\n
Mihaela P
ertea is an Associate Professor in the Department of Biomedical Engineerin
g at Johns Hopkins University. She received her B.S. and M.S. degrees in C
omputer Science from University of Bucharest in Romania\, and her Ph.D in
Computer Science from the Johns Hopkins University School of Engineering.
Dr. Pertea’s work in computational biology draws upon techniques and data
from multiple disciplines\, including computer science and molecular biolo
gy\, genetics\, biotechnology\, and statistics. Her work has focused on co
mputational gene finding and sequence pattern recognition and she has deve
loped several open-source gene finders that were used for the annotation o
f the genomes of Plasmodium falciparum (malaria parasite)\, A
rabidopsis thaliana\, rice\, Aspergillus fumigatus\, Cryptococcus
neoformans\, and others. A major focus of her current research is on
developing innovative and efficient methods to analyze large DNA and RNA
sequence data in order to provide a genome-scale understanding of cellular
function. Dr. Pertea believes that the principled use of algorithms from
other fields\, adapted to the problems of computational biology and couple
d with careful software engineering and high performance computing\, has t
he potential to make a significant impact in the life sciences. She has pu
blished over 50 scientific papers that have received more than 30\,000 cit
ations to date.
\n
\n
To join the live event please request th
e link by emailing: icm@jhu.edu
\n
\n
▶RECORDING
\n
\n
<
h4 class='visible-phone visible-xs'>\n Abstract \n
\n<
h2 style='text-align: center\;'>
“Towards a New Comprehensive Human Gene Catalogue<
/span>”\n\n
\n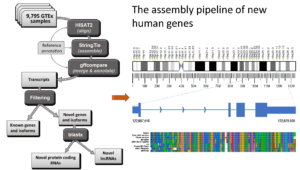
\n
\n
\nA hu
ge and still-growing number of genetic studies depend on the human gene ca
talogue\, including thousands of experiments each year and an enormous inv
estment of time and effort. However\, despite its critical role in biomedi
cal research\, the human gene list is still incomplete and\, in many ways\
, unstable. The widespread use of RNA sequencing technology over the past
decade has allowed scientists to discover a far larger and richer repertoi
re of genes and transcripts than previously known. Our own recent efforts
led to the creation of a new human gene catalogue\, called CHESS\, that w
e built using a very large collection of nearly 10\,000 RNA-seq experiment
s from 31 tissues\, all sequenced as part of the GTEx project. Processing
this large amount of data was one of the most challenging tasks\, made pos
sible by the computational efficiency of StringTie\, a transcriptome assem
bler we developed in our lab.
\n
\n▶RE
CORDING
\n
\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984268@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series\,Events
CONTACT:Mishka Colombo\; 4105164116\; mishka@jhu.edu\; email mishka@jhu.edu
for link
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Exploring the Potentials of Selective Blocking of the IL-1 Syste
m by Distinct Targeting of the Co-receptor TILRR”\n\nDr. Qwarnstrom’s rese
arch focuses on regulation of receptor function and cell signalling. Work
on signal transduction\, has centred on regulation of NF-B pathways and o
n using single cell recordings of regulatory events in live cells. The sin
gle cell data was used as the basis for developing highly detailed\, predi
ctive models of the IL-1 system and the NF-B network. Interdisciplinary p
rojects led to identification and characterisation of the IL-1R1 co-recept
or TILRR and established its role in IL-1 receptor function\, NF-B regula
tion and disease. Ongoing work focuses on evaluating TILRR as a potential
therapeutic target.\n \nTo join the live event please request the link by
emailing: icm@jhu.edu\n \n▶RECORDING \n\n \n Abstract \n\n“
Exploring the Potentials of Selective Blocking of the IL-1 System by Disti
nct Targeting of the Co-receptor TILRR”\n \n\n \n \nAberrant activation of
NF-B plays a central role in disease. IL-1 is a key regulator of NF-B a
nd has emerged as a rational therapeutic target. Clinical trials have demo
nstrated that nonselective blocking of the IL-1 system can cause serious s
ide effects related to impairment of the immune system\, highlighting the
need for more specific targeting. Our interdisciplinary studies on regulat
ion of NF-B led to identification of the IL1R1 coreceptor TILRR and estab
lished its role in controlling IL1R1 function and in driving aberrant acti
vation of NF-B. Our published work shows that genetic deletion or antibod
y blocking of TILRR reduces progression of inflammatory conditions. Pathwa
y enrichment analysis of TILRR-induced gene expression profiles has reveal
ed significant links with NF-B signalling\, Alzheimer’s disease and cance
r. I will describe the predictive modelling approaches used in these studi
es\, outline the key regulatory events that underpin TILRR’s control of th
e IL-1 system and selective regulation the NF-B pathways\, present recent
data on the role of TILRR in disease and discuss its potential as a thera
peutic target.\n \n \n▶RECORDING
DTSTART;TZID=America/New_York:20220503T103000
DTEND;TZID=America/New_York:20220503T113000
LOCATION:Zoom
SEQUENCE:0
SUMMARY:Exploring the Potentials of Selective Blocking of the IL-1 System b
y Distinct Targeting of the Co-receptor TILRR
URL:https://icm.jhu.edu/events/exploring-the-potentials-of-selective-blocki
ng-of-the-il-1-system-by-distinct-targeting-of-the-co-receptor-tilrr/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2022/03/E
Qwarnstrom-e1648747764368.jpg\;163\;183\,medium\;https://icm.jhu.edu/wp-co
ntent/uploads/2022/03/EQwarnstrom-e1648747764368.jpg\;163\;183\,large\;htt
ps://icm.jhu.edu/wp-content/uploads/2022/03/EQwarnstrom-e1648747764368.jpg
\;163\;183\,full\;https://icm.jhu.edu/wp-content/uploads/2022/03/EQwarnstr
om-e1648747764368.jpg\;163\;183
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Explor
ing the Potentials of Selective Blocking of the IL-1 System by Distinct Ta
rgeting of the Co-receptor TILRR”
\n

\n
Dr. Qwarnstro
m’s research focuses on regulation of receptor function and cell signallin
g. Work on signal transduction\, has centred on regulation of NF-B pathwa
ys and on using single cell recordings of regulatory events in live cells.
The single cell data was used as the basis for developing highly detailed
\, predictive models of the IL-1 system and the NF-B network. Interdiscip
linary projects led to identification and characterisation of the IL-1R1 c
o-receptor TILRR and established its role in IL-1 receptor function\, NF-
B regulation and disease. Ongoing work focuses on evaluating TILRR as a po
tential therapeutic target.
\n
\n
To join the live event pleas
e request the link by emailing: icm@j
hu.edu
\n
\n
▶RECORDING
\n
\n
\n Abstract
\n
\n
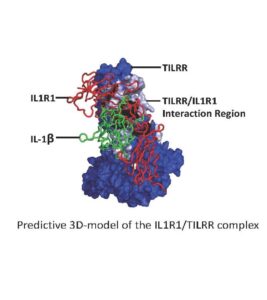
“Exploring the Potentials of Selective Blocking of
the IL-1 System by Distinct Targeting of the Co-receptor TILRR”
\n
\n
\n
\n
\n
Aberrant activation of NF-B plays a central role in disease. IL
-1 is a key regulator of NF-B and has emerged as a rational therapeutic t
arget. Clinical trials have demonstrated that nonselective blocking of the
IL-1 system can cause serious side effects related to impairment of the i
mmune system\, highlighting the need for more specific targeting. Our inte
rdisciplinary studies on regulation of NF-B led to identification of the
IL1R1 coreceptor TILRR and established its role in controlling IL1R1 funct
ion and in driving aberrant activation of NF-B. Our published work shows
that genetic deletion or antibody blocking of TILRR reduces progression of
inflammatory conditions. Pathway enrichment analysis of TILRR-induced gen
e expression profiles has revealed significant links with NF-B signalling
\, Alzheimer’s disease and cancer. I will describe the predictive modellin
g approaches used in these studies\, outline the key regulatory events tha
t underpin TILRR’s control of the IL-1 system and selective regulation the
NF-B pathways\, present recent data on the role of TILRR in disease and
discuss its potential as a therapeutic target.
\n
\n
\n
▶RECORDING
\n
\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984426@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series
CONTACT:
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Elucidating the Neural Control of Movement Using AI”\n\nShreya S
axena is broadly interested in the neural control of complex\, coordinated
behavior. She is an Assistant Professor at the University of Florida’s De
partment of Electrical and Computer Engineering. Before this\, Shreya was
a Swiss National Science Foundation Postdoctoral Fellow at Columbia Univer
sity’s Zuckerman Mind Brain Behavior Institute. She did her PhD in the Dep
artment of Electrical Engineering and Computer Science at the Massachusett
s Institute of Technology studying the closed-loop control of fast movemen
ts from a control theory perspective. Shreya received a B.S. in Mechanical
Engineering from the Swiss Federal Institute of Technology (EPFL)\, and a
n M.S. in Biomedical Engineering from Johns Hopkins University. She is hon
ored to have been selected as a Rising Star in both Electrical Engineering
(2019) and Biomedical Engineering (2018).\n \n▶RECORDING \n\n \n
Abstract \n\n“Elucidating the Neural Control of Movement Using AI”
\n \n\n \n \n \nHow does the motor cortex achieve generalizable and purpos
eful movements from the complex\, nonlinear musculoskeletal system? Prev
ious research in the field typically does not consider the biophysical und
erpinnings of the musculoskeletal system\, and thus fails to elucidate the
computational role of neural activity in driving the musculoskeletal syst
em such that the body reaches a desired state. Here\, I will present a dee
p reinforcement learning framework for training recurrent neural network c
ontrollers that act on anatomically accurate limb models such that they ac
hieve desired movements. We apply this framework to kinematic and neural r
ecordings made in macaques as they perform movements at different speeds.
This framework for the control of the musculoskeletal system mimics biolog
ically observed neural strategies\, and enables hypothesis generation for
prediction and analysis of novel movements and neural strategies.\nEffecti
vely modeling and quantifying behavior is essential for our understanding
of the brain. Modeling behavior across different subjects and in a natural
istic setting remains a significant challenge in the field of behavioral q
uantification. We develop novel explainable AI methods for modeling contin
uously varying differences in behavior\, which successfully represent dist
inct features of multi-subject and social behavior in an unsupervised mann
er. These methods are also successful at uncovering the relationships betw
een recorded neural data and the ensuing behavior. I will end with future
avenues on explainable AI methods for elucidating the neural control of mo
vement.\n \n▶RECORDING
DTSTART;TZID=America/New_York:20220906T103000
DTEND;TZID=America/New_York:20220906T113000
LOCATION:Levering Hall: Great Hall
SEQUENCE:0
SUMMARY:Elucidating the Neural Control of Movement Using AI
URL:https://icm.jhu.edu/events/elucidating-the-neural-control-of-movement-u
sing-ai/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2022/08/S
hreya-291x300.jpeg\;183\;189\,medium\;https://icm.jhu.edu/wp-content/uploa
ds/2022/08/Shreya-291x300.jpeg\;183\;189\,large\;https://icm.jhu.edu/wp-co
ntent/uploads/2022/08/Shreya-291x300.jpeg\;183\;189\,full\;https://icm.jhu
.edu/wp-content/uploads/2022/08/Shreya-291x300.jpeg\;183\;189
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Elucid
ating the Neural Control of Movement Using AI”
\n

\n
Shreya Saxena is broadly interested
in the neural control of complex\, coordinated behavior. She is an Assist
ant Professor at the University of Florida’s Department of Electrical and
Computer Engineering. Before this\, Shreya was a Swiss National Science Fo
undation Postdoctoral Fellow at Columbia University’s Zuckerman Mind Brain
Behavior Institute. She did her PhD in the Department of Electrical Engin
eering and Computer Science at the Massachusetts Institute of Technology s
tudying the closed-loop control of fast movements from a control theory pe
rspective. Shreya received a B.S. in Mechanical Engineering from the Swiss
Federal Institute of Technology (EPFL)\, and an M.S. in Biomedical Engine
ering from Johns Hopkins University. She is honored to have been selected
as a Rising Star in both Electrical Engineering (2019) and Biomedical Engi
neering (2018).
\n
\n
<
a style='color: #993300\;' href='https://wse.zoom.us/rec/share/25b21s-2e4H
aHownnYYH7ggruOxn_5KUSGHLzZ1TBawOKb5ypWQSaVczCg6S76LC.vrlHqfjcd8jGWLaU?sta
rtTime=1662473228000' target='_blank' rel='noopener'>▶RECORDING
\n
\n
\n Abstract
\n
\n
“Elucidating the Neural C
ontrol of Movement Using AI
”
\n
\n

\n
\n
\n
\n
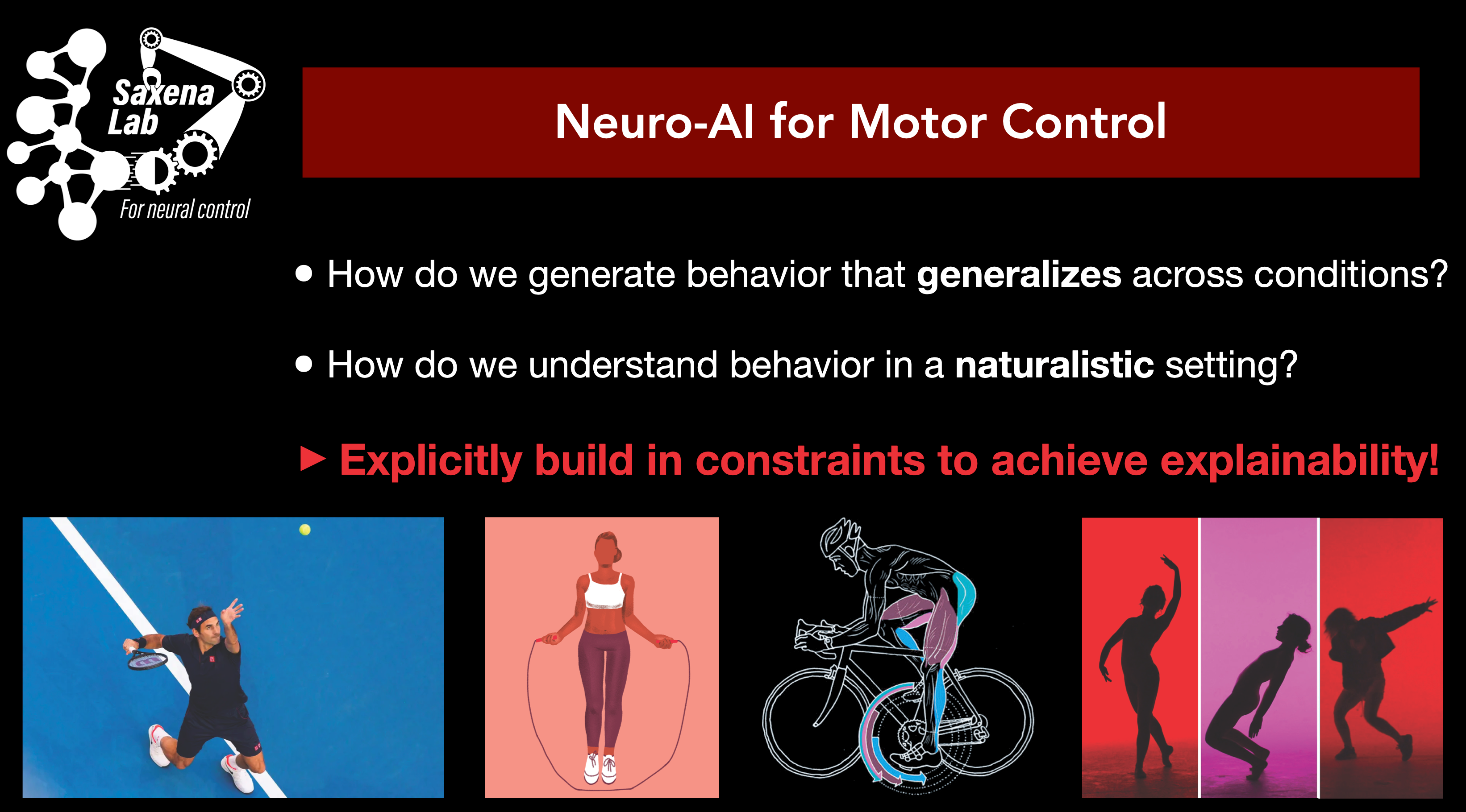
How does the motor cortex achieve generalizable and purpo
seful movements from the complex\, nonlinear musculoskeletal system? Pre
vious research in the field typically does not consider the biophysical un
derpinnings of the musculoskeletal system\, and thus fails to elucidate th
e computational role of neural activity in driving the musculoskeletal sys
tem such that the body reaches a desired state. Here\, I will present a de
ep reinforcement learning framework for training recurrent neural network
controllers that act on anatomically accurate limb models such that they a
chieve desired movements. We apply this framework to kinematic and neural
recordings made in macaques as they perform movements at different speeds.
This framework for the control of the musculoskeletal system mimics biolo
gically observed neural strategies\, and enables hypothesis generation for
prediction and analysis of novel movements and neural strategies.
\n
Effectively modeling and quantifying behavior is essential for our unders
tanding of the brain. Modeling behavior across different subjects and in a
naturalistic setting remains a significant challenge in the field of beha
vioral quantification. We develop novel explainable AI methods for modelin
g continuously varying differences in behavior\, which successfully repres
ent distinct features of multi-subject and social behavior in an unsupervi
sed manner. These methods are also successful at uncovering the relationsh
ips between recorded neural data and the ensuing behavior. I will end with
future avenues on explainable AI methods for elucidating the neural contr
ol of movement.
\n
\n
▶RECORDING
\n
\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984560@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series\,Events
CONTACT:Mishka Colombo\; 410-516-4116\; mcolomb4@jhu.edu\; https://tinyurl.
com/2tcz6w4s
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
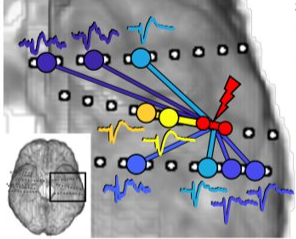
Bio \n“Significance of Event Related Causality (ERC) in Neural Networks
”\n\nDr. Anna Korzeniewska obtained MS in Physics with concentration in Me
dical Physics from University of Warsaw\, Poland and PhD in Biological Sci
ences with concentration in Neurophysiology from Nencki Institute of Exper
imental Biology\, Polish Academy of Sciences. Since 2004 she works at John
s Hopkins’ Epilepsy Center. Her research interest is focused on the dynami
cs of causal interactions among functional and pathological neural network
s.\n \n▶RECORDING\n\n \n Abstract \n\n\n“Significance of Ev
ent Related Causality (ERC) in Neural Networks”\n\n\n \nNeural activity is
propagated across large-scale cortical networks on very brief time scales
. Studying such transient and complex systems calls for a short time-windo
w on one hand\, and a great extent of recording sites in the brain\, on th
e other. These demands are not easily satisfied\, as short time intervals
do not provide enough data-points to model the dynamics of large-scale bra
in networks. The limitation can be overcome by using multiple realizations
of the same process\, but the price to be paid is that traditional statis
tical methods cannot be used to assess the significance of event-related c
hanges in the estimated dynamics of the system. To obtain statistical conf
idence of the dynamics of neural interactions among large-scale networks r
evealed by event-related causality (ERC)\, we propose using the variance o
f a two-dimensional moving average. We also propose a criterion for the tw
o-dimensional model selection\, which combines the difference between the
smooth estimator and the real values with the confidence interval. We show
that this estimator is efficient\, stable\, and ensures precise embedding
of statistical significance in two-dimensional (time-frequency) space. He
re\, we show that the method can be used to investigate information flow a
mong eloquent network\, to provide a guidance for epileptic surgery.\n \n▶
RECORDING
DTSTART;TZID=America/New_York:20221004T103000
DTEND;TZID=America/New_York:20221004T113000
LOCATION:Levering Hall: Great Hall @ 3400 N. Charles Street
SEQUENCE:0
SUMMARY:Significance of Event Related Causality (ERC) in Neural Networks
URL:https://icm.jhu.edu/events/significance-of-event-related-causality-erc-
in-neural-networks/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2022/09/J
HUanna_small-290x300.jpg\;222\;230\,medium\;https://icm.jhu.edu/wp-content
/uploads/2022/09/JHUanna_small-290x300.jpg\;222\;230\,large\;https://icm.j
hu.edu/wp-content/uploads/2022/09/JHUanna_small-290x300.jpg\;222\;230\,ful
l\;https://icm.jhu.edu/wp-content/uploads/2022/09/JHUanna_small-290x300.jp
g\;222\;230
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Signif
icance of Event Related Causality (ERC) in Neural Networks
”
\n

\n
Dr. Anna Korzeniewska obtained MS in Physics with concentration
in Medical Physics from University of Warsaw\, Poland and PhD in Biologic
al Sciences with concentration in Neurophysiology from Nencki Institute of
Experimental Biology\, Polish Academy of Sciences. Since 2004 she works a
t Johns Hopkins’ Epilepsy Center. Her research interest is focused on the
dynamics of causal interactions among functional and pathological neural n
etworks.
\n
\n
▶RECORDING
\n
\n
\n Abstract
\n
\n
\n
“
Significance of Event Related Causality (ERC) in Neural Networks<
/span>”
\n
\n
 \n
\n
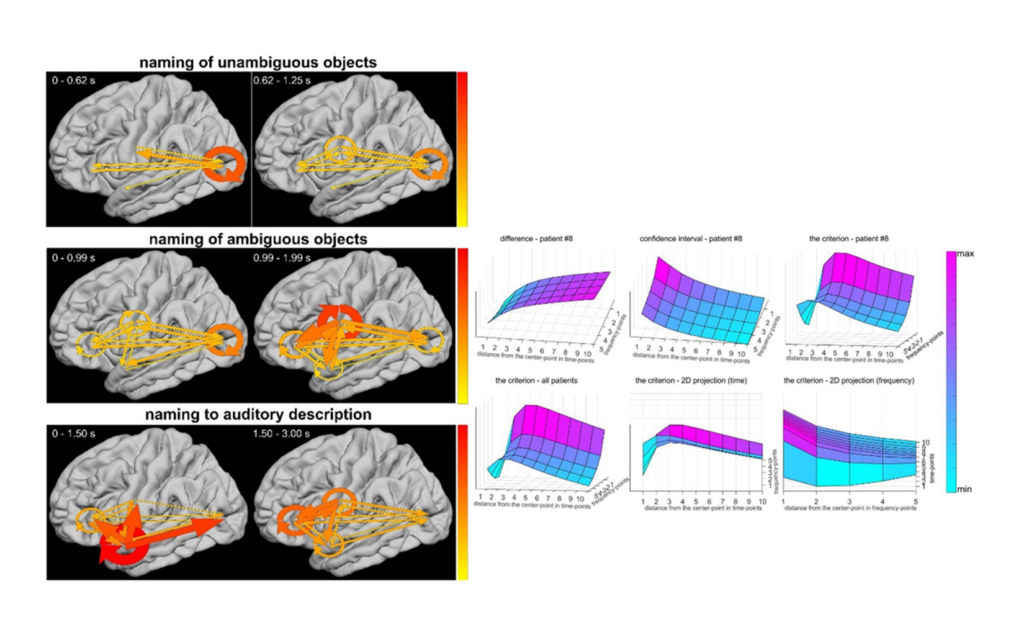
\n
Neural activity is propagated across large-scale cortical
networks on very brief time scales. Studying such transient and complex s
ystems calls for a short time-window on one hand\, and a great extent of r
ecording sites in the brain\, on the other. These demands are not easily s
atisfied\, as short time intervals do not provide enough data-points to mo
del the dynamics of large-scale brain networks. The limitation can be over
come by using multiple realizations of the same process\, but the price to
be paid is that traditional statistical methods cannot be used to assess
the significance of event-related changes in the estimated dynamics of the
system. To obtain statistical confidence of the dynamics of neural intera
ctions among large-scale networks revealed by event-related causality (ERC
)\, we propose using the variance of a two-dimensional moving average. We
also propose a criterion for the two-dimensional model selection\, which c
ombines the difference between the smooth estimator and the real values wi
th the confidence interval. We show that this estimator is efficient\, sta
ble\, and ensures precise embedding of statistical significance in two-dim
ensional (time-frequency) space. Here\, we show that the method can be use
d to investigate information flow among eloquent network\, to provide a gu
idance for epileptic surgery.
\n
\n
▶RECOR
DING
\n
\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984606@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES:
CONTACT:Mishka\; 14105164116\; mcolomb4@jhu.edu\; https://tinyurl.com/3w5cc
3er
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Biophysically Realistic Cortical Network Models for Simulation o
f Cortical and Intracortical Electrical Stimulations”\n\nDr. Kudela is an
expert in neural signal data modeling and analysis. His research is focuse
d on computational modeling of cortical dynamics (cortical electrical stim
ulation\, cortical auditory processing\, and seizure) to rationalize exper
imental observations from novel microelectrode recordings in invasively mo
nitored epilepsy patients. He is a computational neuroscientist\, and skil
led in many domains ranging from computer science and scientific programmi
ng to parallel computing and high-performance computing. He collaborates w
ith several Johns Hopkins investigators working on new medical therapies a
nd devices.\n \n▶RECORDING \n\n \n Abstract \n\n\n“Biophysi
cally Realistic Cortical Network Models for Simulation of Cortical and Int
racortical Electrical Stimulations”\n\n\n \nModeling electrical stimulatio
n of neural elements can be performed in two steps. The first step involve
s the calculation of the spatial distributions of the induced electric fie
lds in cortical volume produced by stimulating electrodes. The second step
is to model the response of neuronal elements to an electric field using
multicompartmental representations of neurons. The response of an individu
al neuron to electrical stimulation is determined by several factors like
neuronal morphology and the cortical geometry that affects electric field
distribution in the cortical volume. We use computational models of cortic
al neurons to investigate the effects of cortical and intracortical electr
ical stimulations in a cortical volume. Two high-resolution cortical netwo
rk models will be presented that were developed to study 1) cortical respo
nses to subdural cortical stimulations and 2) neuronal recruitment by intr
acortical microstimulation for restoring touch sensation.\n \n▶RECORDING
DTSTART;TZID=America/New_York:20221101T103000
DTEND;TZID=America/New_York:20221101T113000
LOCATION:Levering Hall: Great Hall
SEQUENCE:0
SUMMARY:Biophysically Realistic Cortical Network Models for Simulation of C
ortical and Intracortical Electrical Stimulations
URL:https://icm.jhu.edu/events/biophysically-realistic-cortical-network-mod
els-for-simulation-of-cortical-and-intracortical-electrical-stimulations/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2022/10/P
icture1.png\;148\;164\,medium\;https://icm.jhu.edu/wp-content/uploads/2022
/10/Picture1.png\;148\;164\,large\;https://icm.jhu.edu/wp-content/uploads/
2022/10/Picture1.png\;148\;164\,full\;https://icm.jhu.edu/wp-content/uploa
ds/2022/10/Picture1.png\;148\;164
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Biophy
sically Realistic Cortical Network Models for Simulation of Cortical and I
ntracortical Electrical Stimulations”<
/h2>\n

\n
Dr. Kudela is an expert in neural signal data modeling and anal
ysis. His research is focused on computational modeling of cortical dynami
cs (cortical electrical stimulation\, cortical auditory processing\, and s
eizure) to rationalize experimental observations from novel microelectrode
recordings in invasively monitored epilepsy patients. He is a computation
al neuroscientist\, and skilled in many domains ranging from computer scie
nce and scientific programming to parallel computing and high-performance
computing. He collaborates with several Johns Hopkins investigators workin
g on new medical therapies and devices.
\n
\n
▶RECORDING
\n
\n
\n Abstract
\n
\n
\n
“Biophysically Realistic Cortical Network M
odels for Simulation of Cortical and Intracortical Electrical Stimulations
”
\n
\n

\n
\n
Modeling
electrical stimulation of neural elements can be performed in two steps.
The first step involves the calculation of the spatial distributions of th
e induced electric fields in cortical volume produced by stimulating elect
rodes. The second step is to model the response of neuronal elements to an
electric field using multicompartmental representations of neurons. The r
esponse of an individual neuron to electrical stimulation is determined by
several factors like neuronal morphology and the cortical geometry that a
ffects electric field distribution in the cortical volume. We use computat
ional models of cortical neurons to investigate the effects of cortical an
d intracortical electrical stimulations in a cortical volume. Two high-res
olution cortical network models will be presented that were developed to s
tudy 1) cortical responses to subdural cortical stimulations and 2) neuron
al recruitment by intracortical microstimulation for restoring touch sensa
tion.
\n
\n
▶RECORDING
\n
\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984668@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES:
CONTACT:Mishka\; 14105164116\; mcolomb4@jhu.edu\; https://icm.jhu.edu/semin
ar-series/
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Evolutionary Dynamics of Tumor Progression”\n\nDr. Bozic is an A
ssociate Professor in the Department of Applied Mathematics at the Univers
ity of Washington. She develops mathematical and computational models to s
tudy the evolutionary dynamics of cancer. Her research interests include m
athematical biology\, stochastic processes and analysis of genomic and cli
nical data.\n \n▶RECORDING \n\n \n Abstract \n\n\n“Evolutio
nary Dynamics of Tumor Progression”\n\n\nCancer is the result of a stochas
tic evolutionary process characterized by the accumulation of mutations th
at are responsible for tumor initiation\, progression\, immune escape\, an
d drug resistance\, as well as mutations with no effect on the phenotype.
Mathematical modeling can be used to describe the dynamics of tumor cell p
opulations and to obtain insights into the hidden evolutionary processes l
eading to cancer. I will present recent approaches that employ stochastic
models of cancer evolution to quantify evolutionary dynamics of chronic ly
mphocytic leukemia and colorectal cancer in patients\, and their implicati
ons for interpretation of cancer sequencing data.\n \n▶RECORDING
DTSTART;TZID=America/New_York:20221206T103000
DTEND;TZID=America/New_York:20221206T113000
LOCATION:Zoom: email mishka@jhu.edu for link
SEQUENCE:0
SUMMARY:“Evolutionary Dynamics of Tumor Progression”
URL:https://icm.jhu.edu/events/evolutionary-dynamics-of-tumor-progression/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2022/11/i
vana-1.jpg\;200\;194\,medium\;https://icm.jhu.edu/wp-content/uploads/2022/
11/ivana-1.jpg\;200\;194\,large\;https://icm.jhu.edu/wp-content/uploads/20
22/11/ivana-1.jpg\;200\;194\,full\;https://icm.jhu.edu/wp-content/uploads/
2022/11/ivana-1.jpg\;200\;194
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Evolut
ionary Dynamics of Tumor Progression”<
/h2>\n

\n
Dr. Bozic is an Associat
e Professor in the Department of Applied Mathematic
s at the University of Washington. Sh
e develops mathematical and computational models to study the evolutionary
dynamics of cancer. Her research interests include mathematical biology\,
stochastic processes and analysis of genomic and clinical data.
\n
\n
▶RECO
RDING
\n
\n
\n Abstract
\n
\n
<
/h2>\n“Evolutionary Dynamics of Tumor Progression<
/span>”
\n\n
\n
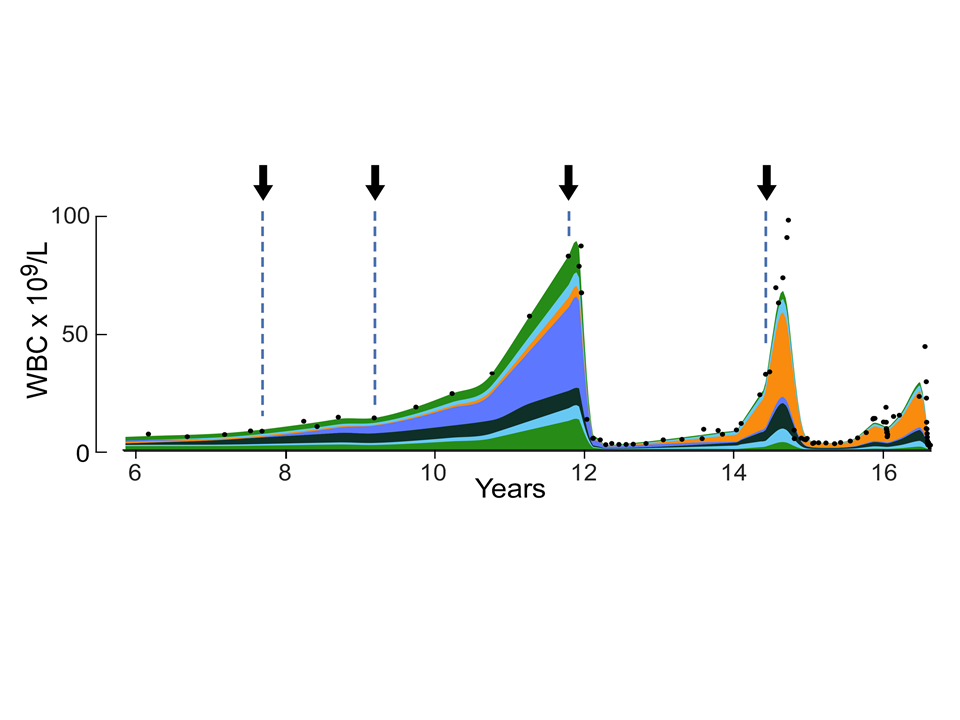
Cancer is the result of a stochastic evolutionary process
characterized by the accumulation of mutations that are responsible for tu
mor initiation\, progression\, immune escape\, and drug resistance\, as we
ll as mutations with no effect on the phenotype. Mathematical modeling can
be used to describe the dynamics of tumor cell populations and to obtain
insights into the hidden evolutionary processes leading to cancer. I will
present recent approaches that employ stochastic models of cancer evolutio
n to quantify evolutionary dynamics of chronic lymphocytic leukemia and co
lorectal cancer in patients\, and their implications for interpretation of
cancer sequencing data.
\n
\n
▶RECORDING
\n
\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984738@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series\,Events\,Special Sem
inars
CONTACT:Mishka Colombo\; 4105164116\; mcolomb4@jhu.edu\; https://tinyurl.co
m/4v8n6rcw
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Resting State fMRI in Epilepsy for Seizure Onset Localization: E
vidence and Methods”\n\nVarina L. Boerwinkle\, MD is the Division Chief of
Child Neurology at the University of North Carolina in Chapel Hill\, and
Professor of Neurology. She is also the medical director of the Functional
Neuroimaging and Neuroscience Laboratory and Pediatric Neurocritical Care
Service. She earned her medical degree from University of Texas Southwest
ern and completed a residency in child neurology at Baylor College of Medi
cine.\nHer clinical and research efforts in brain networks began in 2010.
She pioneered the clinical utilization of resting state functional MRI (rs
-fMRI) for children to localize seizure onset zones\, and brain networks\,
and improve epilepsy surgery outcomes. Through her efforts\, over 2000 in
dividual children primarily with epilepsy have received rs-fMRI with clini
cally impactful results. For many of these children\, who are unable to pe
rform demanding tests reliably\, rs-fMRI\, analyzed by methods validated i
n her lab\, offers comprehensive major brain network characterization with
the capacity for clinical correlation.\n \n▶RECORDING \n\n \n
Abstract \n\n\n“Resting State fMRI in Epilepsy for Seizure Onset Loca
lization: Evidence and Methods”\n\n\nEpilepsy effects over 50 million worl
dwide and the only known cure is surgery. However\, the success of the sur
gery relies on accurate localization of the seizure onset zone\, which wit
h standard techniques ranges 30-70%. Recently\, resting state fMRI has bee
n shown to not only show the normal networks to avoid surgical morbidity\,
but to also localize the seizure network. In this presentation we will re
view the evidence behind this new diagnostic and discuss potential avenue
for future investigations. .\n \n▶RECORDING
DTSTART;TZID=America/New_York:20230207T103000
DTEND;TZID=America/New_York:20230207T113000
LOCATION:Levering Hall: Great Hall
SEQUENCE:0
SUMMARY:Resting State fMRI in Epilepsy for Seizure Onset Localization: Evi
dence and Methods
URL:https://icm.jhu.edu/events/resting-state-fmri-in-epilepsy-for-seizure-o
nset-localization-evidence-and-methods/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2023/01/B
oerwinkle-scaled-e1673628465991.jpg\;158\;168\,medium\;https://icm.jhu.edu
/wp-content/uploads/2023/01/Boerwinkle-scaled-e1673628465991.jpg\;158\;168
\,large\;https://icm.jhu.edu/wp-content/uploads/2023/01/Boerwinkle-scaled-
e1673628465991.jpg\;158\;168\,full\;https://icm.jhu.edu/wp-content/uploads
/2023/01/Boerwinkle-scaled-e1673628465991.jpg\;158\;168
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Restin
g State fMRI in Epilepsy for Seizure Onset Localization: Evidence and Meth
ods”
\n
 \n
\n
Varina L. Boerwinkle\, MD is the Division Chief of Child Neurology
at the University of North Carolina in Chapel Hill\, and Professor of Neur
ology. She is also the medical director of the Functional Neuroimaging and
Neuroscience Laboratory and Pediatric Neurocritical Care Service. She ear
ned her medical degree from University of Texas Southwestern and completed
a residency in child neurology at Baylor College of Medicine.
\n
Her
clinical and research efforts in brain networks began in 2010. She pionee
red the clinical utilization of resting state functional MRI (rs-fMRI) for
children to localize seizure onset zones\, and brain networks\, and impro
ve epilepsy surgery outcomes. Through her efforts\, over 2000 individual c
hildren primarily with epilepsy have received rs-fMRI with clinically impa
ctful results. For many of these children\, who are unable to perform dema
nding tests reliably\, rs-fMRI\, analyzed by methods validated in her lab\
, offers comprehensive major brain network characterization with the capac
ity for clinical correlation.
\n
\n
▶RECOR
DING
\n
\n
\n Abstract
\n
\n
<
/h2>\n“Resting State fMRI in Epilepsy for Seizure Onset Lo
calization: Evidence and Methods”
\n\n
 <
/a>
<
/a>
\n
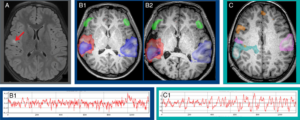
Epilepsy effects over 50 million worldwide and the only known
cure is surgery. However\, the success of the surgery relies on accurate l
ocalization of the seizure onset zone\, which with standard techniques ran
ges 30-70%. Recently\, resting state fMRI has been shown to not only show
the normal networks to avoid surgical morbidity\, but to also localize the
seizure network. In this presentation we will review the evidence behind
this new diagnostic and discuss potential avenue for future investigations
. .
\n
\n
▶RECORDING
\n\n
\n
\n \n
X-TAGS;LANGUAGE=en-US:Distinguished Seminar Series\,Varina Boerwinkle
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984763@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series
CONTACT:Mishka Colombo\; 4105164116\; mcolomb4@jhu.edu\; https://wse.zoom.u
s/j/92808587799
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Harnessing Artificial\nIntelligence for Healthcare”\n\n \nSushmi
ta Mitra is a full professor at the Machine Intelligence Unit (MIU)\, Indi
an Statistical Institute\, Kolkata. From 1992 to 1994 she was in the RWTH\
, Aachen\, Germany as a DAAD Fellow. She was a Visiting Professor in the C
omputer Science Departments of the University of Alberta\, Edmonton\, Cana
da\; Meiji University\, Japan\; and Aalborg University Esbjerg\, Denmark.
Dr. Mitra received the National Talent Search Scholarship (1978-1983) from
NCERT\, India\, the University Gold Medal in 1988\, the IEEE TNN Outstand
ing Paper Award in 1994 for her pioneering work in neuro-fuzzy computing\,
the CIMPA-INRIA-UNESCO Fellowship in 1996\, and Fulbright-Nehru Senior Re
search Fellowship in 2018-2020. She was the INAE Chair Professor during 20
18-2020. Dr. Mitra has been awarded the prestigious J. C. Bose National Fe
llowship\, 2021.\nDr. Mitra is the author of the books “Neuro-Fuzzy Patter
n Recognition: Methods in Soft Computing” and “Data Mining: Multimedia\, S
oft Computing\, and Bioinformatics” published by John Wiley\, and “Introdu
ction to Machine Learning and Bioinformatics”\, Chapman & Hall/CRC Press\,
beside a host of other edited books. Dr. Mitra has guest edited special i
ssues of several journals\, is an Associate Editor of “IEEE/ACM Trans. on
Computational Biology and Bioinformatics“\, “Information Sciences“\, “Fund
amenta Informatica“\, “Computers in Biology and Medicine“\, SN Computer Sc
iences and is a Founding Associate Editor of “Wiley Interdisciplinary Revi
ews: Data Mining and Knowledge Discovery (WIRE DMKD)“. She has more than 1
50 research publications in referred international journals. According to
the Stanford List\, Dr. Mitra is ranked among the top 2% scientists worldw
ide in the domain of Artificial Intelligence and Image Processing.\nDr. Mi
tra is a Fellow of the IEEE\, The World Academy of Sciences (TWAS)\, India
n National Science Academy (INSA)\, International Association for Pattern
Recognition (IAPR)\, Asia-Pacific Artificial Intelligence Association (AAI
A)\, and Fellow of the Indian National Academy of Engineering (INAE) and T
he National Academy of Sciences\, India (NASI). She serves as a Member of
the Inter-Academy Panel Panel for Women in STEMM. She has visited more tha
n 30 countries as a Plenary/Invited Speaker or an academic visitor. She se
rved in the capacity of General Chair\, Program Chair\, Tutorial Chair\, o
f many international conferences\; was the Chair\, IEEE Kolkata Section (2
021-2022) and an IEEE CIS Distinguished Lecturer. Her current research int
erests include data science\, machine learning\, soft computing\, medical
image processing\, and Bioinformatics.\n \n▶RECORDING \n\n \n
Abstract \n\n\n“Harnessing Artificial\nIntelligence for Healthcare”\n
\nThe talk will focus on the role of Artificial Intelligence and Learning
in the domain of healthcare. Topics like Genomics\, Radiomics\, Radiogenom
ics\, and Personalized Medicine will be discussed in this perspective. Som
e research applications made by our group in these areas will be described
. These include segmentation and survival prediction in GBM tumors from MR
I scans of the brain\; screening of covid -19 from X-ray images of the lun
gs\; and early detection of diabetic retinopathy from fundus images of the
eye.\n \n▶RECORDING
DTSTART;TZID=America/New_York:20230307T103000
DTEND;TZID=America/New_York:20230307T113000
LOCATION:Gilman Hall: Gilman 50 (Marjorie Fisher Auditorium)
SEQUENCE:0
SUMMARY:Harnessing Artificial Intelligence for Healthcare
URL:https://icm.jhu.edu/events/harnessing-artificial-intelligence-for-healt
hcare/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2023/02/S
hushmita-Mitra.jpg\;213\;200\,medium\;https://icm.jhu.edu/wp-content/uploa
ds/2023/02/Shushmita-Mitra.jpg\;213\;200\,large\;https://icm.jhu.edu/wp-co
ntent/uploads/2023/02/Shushmita-Mitra.jpg\;213\;200\,full\;https://icm.jhu
.edu/wp-content/uploads/2023/02/Shushmita-Mitra.jpg\;213\;200
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Harnes
sing Artificial
\nIntelligence for Healthcare”<
/span>
\n
 <
/p>\n
<
/p>\n
\n
Sushmita Mitra is a full professor at the M
achine Intelligence Unit (MIU)\, Indian Statistical Institute\, Kolkata. F
rom 1992 to 1994 she was in the RWTH\, Aachen\, Germany as a DAAD Fellow.
She was a Visiting Professor in the Computer Science Departments of
the University of Alberta\, Edmonton\, Canada\; Meiji University\, Japan\
; and Aalborg University Esbjerg\, Denmark. Dr. Mitra received the Nationa
l Talent Search Scholarship (1978-1983) from NCERT\, India\, the Univer
sity Gold Medal in 1988\, the IEEE TNN Outstanding Paper Award
in 1994 for her pioneering work in neuro-fuzzy computing\, the CIMPA-INRIA
-UNESCO Fellowship in 1996\, and Fulbright-Nehru Senior Research Fellow
ship in 2018-2020. She was the INAE Chair Professor during 2018
-2020. Dr. Mitra has been awarded the prestigious J. C. Bose National Fell
owship\, 2021.
\n
Dr. Mitra is the author of the books “
Neuro-Fuzzy Pattern Recognition: Methods in Soft Computing” and “Data Mini
ng: Multimedia\, Soft Computing\, and Bioinformatics” published by John Wi
ley\, and “Introduction to Machine Learning and Bioinformatics”\, Chapman
& Hall/CRC Press\, beside a host of other edited books. Dr. Mitra has gues
t edited special issues of several journals\, is an Associate Editor of “<
i>IEEE/ACM Trans. on Computational Biology and Bioinformatics“\, “I
nformation Sciences“\, “Fundamenta Informatica“\, “Computers
in Biology and Medicine“\, SN Computer Sciences and is a Found
ing Associate Editor of “Wiley Interdisciplinary Reviews: Data Mining a
nd Knowledge Discovery (WIRE DMKD)“. She has more than 150 research pu
blications in referred international journals. According to the Stanford L
ist\, Dr. Mitra is ranked among the top 2% scientists worldwide in the dom
ain of Artificial Intelligence and Image Processing.
\n
Dr. Mitra is a Fellow of the IEEE\, The World Academy of Sciences (
TWAS)\, Indian National Science Academy (INSA)\, International Association
for Pattern Recognition (IAPR)\, Asia-Pacific Artificial Intelligence Ass
ociation (AAIA)\, and Fellow of the Indian National Academy of Engi
neering (INAE) and The National Academy of Sciences\, India (NASI). She se
rves as a Member of the Inter-Academy Panel Panel for Women in STEMM. She
has visited more than 30 countries as a Plenary/Invited Speaker or an acad
emic visitor. She served in the capacity of General Chair\, Program Chair\
, Tutorial Chair\, of many international conferences\; was the Chair\, IEE
E Kolkata Section (2021-2022) and an IEEE CIS Distinguished Lecturer. Her current research interests include data science\, machine learning\
, soft computing\, medical image processing\, and Bioinformatics.
\n
\n
▶RECORDING
\n
\n
\n Abstract
\n
\n
\n
“Harnessing Artificial
\nIntelligence for Healthcare”<
/strong>
\n
\n
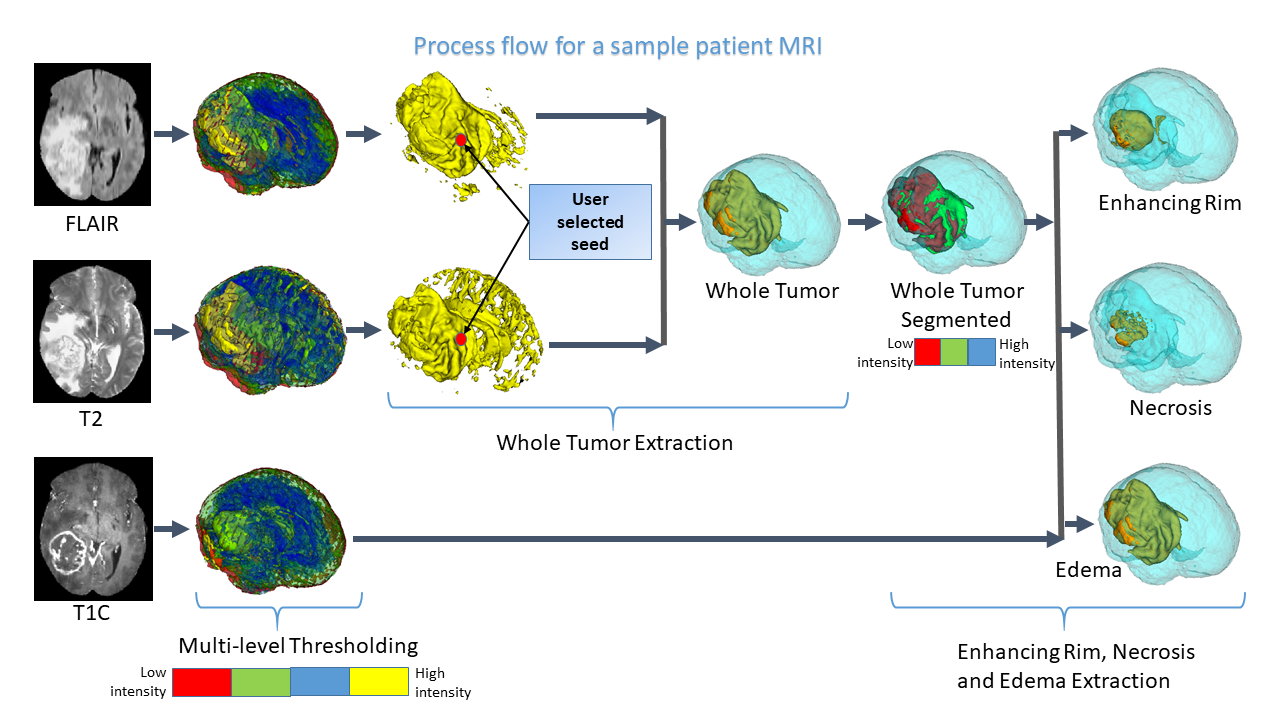 The
talk will focus on the role of Artificial Intelligence and Learning in the
domain of healthcare. Topics like Genomics\, Radiomics\, Radiogenomics\,
and Personalized Medicine will be discussed in this perspective. Some rese
arch applications made by our group in these areas will be described. Thes
e include segmentation and survival prediction in GBM tumors from MRI scan
s of the brain\; screening of covid -19 from X-ray images of the lungs\; a
nd early detection of diabetic retinopathy from fundus images of the eye.<
/p>\n
The
talk will focus on the role of Artificial Intelligence and Learning in the
domain of healthcare. Topics like Genomics\, Radiomics\, Radiogenomics\,
and Personalized Medicine will be discussed in this perspective. Some rese
arch applications made by our group in these areas will be described. Thes
e include segmentation and survival prediction in GBM tumors from MRI scan
s of the brain\; screening of covid -19 from X-ray images of the lungs\; a
nd early detection of diabetic retinopathy from fundus images of the eye.<
/p>\n
\n
▶RECORDING<
/a>
\n
\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984789@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Events\,Special Seminars
CONTACT:Sabrina Sengupta\; ssengu19@jhu.edu
DESCRIPTION:Register for the event by clicking here.
DTSTART;TZID=America/New_York:20230327T170000
DTEND;TZID=America/New_York:20230327T200000
LOCATION:Hackerman B17
SEQUENCE:0
SUMMARY:CM Night
URL:https://icm.jhu.edu/events/cm-night/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2023/03/C
M-Night-Flyer-2023.png\;8203\;10626\,medium\;https://icm.jhu.edu/wp-conten
t/uploads/2023/03/CM-Night-Flyer-2023.png\;8203\;10626\,large\;https://icm
.jhu.edu/wp-content/uploads/2023/03/CM-Night-Flyer-2023.png\;8203\;10626\,
full\;https://icm.jhu.edu/wp-content/uploads/2023/03/CM-Night-Flyer-2023.p
ng\;8203\;10626
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
Register for
the event by clicking here.
\n
\n
X-TAGS;LANGUAGE=en-US:CM Night
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984874@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series
CONTACT:Mishka Colombo\; 4105164116\; mcolomb4@jhu.edu\; https://wse.zoom.u
s/j/96264419972
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Reverse Engineering Chronic Mechanisms of Closed-Loop Brain Stim
ulation for Neuropsychiatric Disorders”\n\nDr. Ankit Khambhati is a biomed
ical engineer who specializes in the development of computational neurotec
hnology to map and modulate large-scale neural circuits affected by brain
network disorders. He is an Assistant Professional Researcher in the Depar
tment of Neurological Surgery at the University of California\, San Franci
sco. There he leads a research program focused on network neuromodulation
and control\, which integrates network science and control theory with bra
in electrical recordings and implantable devices to identify electrophysio
logic biomarkers and develop stimulation-based strategies for rehabilitati
ng or rewiring impaired circuits. Dr. Khambhati is currently using these t
echniques to investigate and optimize closed-loop brain stimulation therap
y for treatment-resistant epilepsy and depression. He previously earned hi
s B.S. in Electrical and Computer Engineering from Carnegie Mellon Univers
ity and Ph.D. in Bioengineering at the University of Pennsylvania\, and he
completed a postdoctoral fellowship in Neuroengineering at the University
of California\, San Francisco.\n \n▶RECORDING\n\n \n Abstract
\n\n\n“Reverse Engineering Chronic Mechanisms of Closed-Loop Brain Sti
mulation for Neuropsychiatric Disorders”\n\n\nClosed-loop neuromodulation
therapy that detects imminent paroxysmal events and rapidly delivers elect
rical stimulation to the brain using a chronically implanted device is an
emerging treatment for pharmacoresistant neurologic or psychiatric disorde
rs. Calibration of closed-loop therapy involves “expert-in-the-loop” optim
ization of device parameters that specify where\, when\, and how electrica
l stimulation pulses should be delivered to the brain for individual patie
nts. A personalized stimulation strategy involves targeting discrete nodes
specific to an individual’s dysfunctional brain network and triggering th
erapeutic stimulation based on neural biomarkers that encode the unique co
nstellation of an individual’s symptoms. In this talk\, I will present an
idealized model of closed-loop stimulation therapy and its adaptation as a
treatment for brain network disorders such as epilepsy and major depressi
ve disorder. Drawing on a range of tools across engineering and neuroscien
ce disciplines — machine learning\, graph theory\, stimulation-based syste
m identification\, and chronic human intracranial EEG from implanted devic
es – I will identify biomarkers related to naturalistic fluctuation in dis
ease state and characterize effects of neurostimulation on functional brai
n network activity and connectivity. Based on these learnings\, I will pro
pose an alternate mechanism of closed-loop therapeutic efficacy based on b
rain network plasticity and discuss opportunities for next-generation devi
ces.\n \n▶RECORDING
DTSTART;TZID=America/New_York:20230404T103000
DTEND;TZID=America/New_York:20230404T113000
LOCATION:Zoom
SEQUENCE:0
SUMMARY:Reverse Engineering Chronic Mechanisms of Closed-Loop Brain Stimula
tion for Neuropsychiatric Disorders
URL:https://icm.jhu.edu/events/reverse-engineering-chronic-mechanisms-of-cl
osed-loop-brain-stimulation-for-neuropsychiatric-disorders/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2023/03/K
hambhati-Headshot-e1679086948434.jpg\;197\;192\,medium\;https://icm.jhu.ed
u/wp-content/uploads/2023/03/Khambhati-Headshot-e1679086948434.jpg\;197\;1
92\,large\;https://icm.jhu.edu/wp-content/uploads/2023/03/Khambhati-Headsh
ot-e1679086948434.jpg\;197\;192\,full\;https://icm.jhu.edu/wp-content/uplo
ads/2023/03/Khambhati-Headshot-e1679086948434.jpg\;197\;192
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Revers
e Engineering Chronic Mechanisms of Closed-Loop Brain Stimulation for Neur
opsychiatric Disorders”
\n

\n
Dr. Ankit Khambhati is a biomedical engineer who s
pecializes in the development of computational neurotechnology to map and
modulate large-scale neural circuits affected by brain network disorders.
He is an Assistant Professional Researcher in the Department of Neurologic
al Surgery at the University of California\, San Francisco. There he leads
a research program focused on network neuromodulation and control\, which
integrates network science and control theory with brain electrical recor
dings and implantable devices to identify electrophysiologic biomarkers an
d develop stimulation-based strategies for rehabilitating or rewiring impa
ired circuits. Dr. Khambhati is currently using these techniques to invest
igate and optimize closed-loop brain stimulation therapy for treatment-res
istant epilepsy and depression. He previously earned his B.S. in Electrica
l and Computer Engineering from Carnegie Mellon University and Ph.D. in Bi
oengineering at the University of Pennsylvania\, and he completed a postdo
ctoral fellowship in Neuroengineering at the University of California\, Sa
n Francisco.
\n
\n
▶RECORDING
\n
\n
\n
Abstract
\n
\n
\n
“Reverse Engineering Chronic Mec
hanisms of Closed-Loop Brain Stimulation for Neuropsychiatric Disorders”
\n
\n
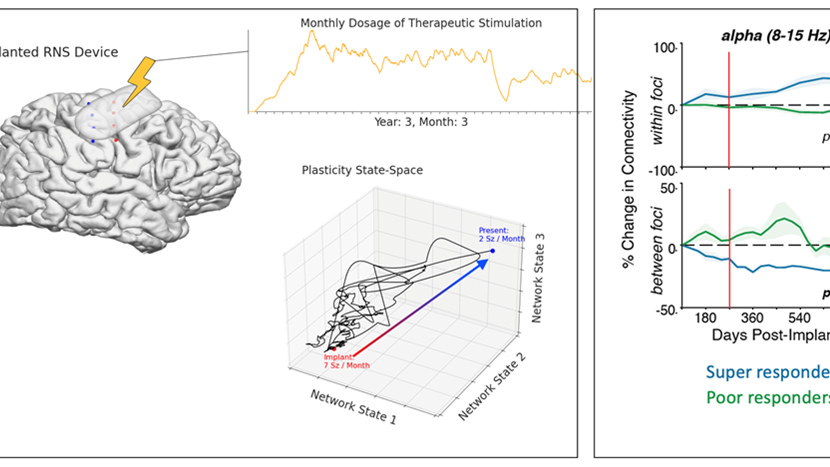
\n
Closed-loop neuromodulation t
herapy that detects imminent paroxysmal events and rapidly delivers electr
ical stimulation to the brain using a chronically implanted device is an e
merging treatment for pharmacoresistant neurologic or psychiatric disorder
s. Calibration of closed-loop therapy involves “expert-in-the-loop” optimi
zation of device parameters that specify where\, when\, and how electrical
stimulation pulses should be delivered to the brain for individual patien
ts. A personalized stimulation strategy involves targeting discrete nodes
specific to an individual’s dysfunctional brain network and triggering the
rapeutic stimulation based on neural biomarkers that encode the unique con
stellation of an individual’s symptoms. In this talk\, I will present an i
dealized model of closed-loop stimulation therapy and its adaptation as a
treatment for brain network disorders such as epilepsy and major depressiv
e disorder. Drawing on a range of tools across engineering and neuroscienc
e disciplines — machine learning\, graph theory\, stimulation-based system
identification\, and chronic human intracranial EEG from implanted device
s – I will identify biomarkers related to naturalistic fluctuation in dise
ase state and characterize effects of neurostimulation on functional brain
network activity and connectivity. Based on these learnings\, I will prop
ose an alternate mechanism of closed-loop therapeutic efficacy based on br
ain network plasticity and discuss opportunities for next-generation devic
es.
\n
\n
▶RECORDING
\n
\n
\n
\n \n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984912@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES:
CONTACT:Mishka Colombo\; 4105164116\; mcolomb4@jhu.edu\; https://wse.zoom.u
s/j/94823269431
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Leveraging Modern Diagnostics to Personalize Treatment for Multi
drug Resistant Tuberculosis”\n\nJeff Tornheim\, MD\, MPH is an Assistant P
rofessor of Medicine\, Pediatrics\, and International Health in the Divisi
on of Infectious Diseases at Johns Hopkins University and a TB clinician a
t the Baltimore City Health Department. He completed a combined residency
in Internal Medicine and Pediatrics at the Yale University School of Medic
ine and a clinical fellowship in Infectious Diseases at the Johns Hopkins
University School of Medicine before joining the faculty in 2017. For the
past 20 years he has engaged in clinical care\, physician education\, and
translational research in India\, South America\, and sub-Saharan Africa.
He is a member of the AIDS Clinical Trials Group Tuberculosis Transformati
ve Science Group\, the IMPAACT Network TB Scientific Committee\, and a pri
ncipal investigator in the RePORT India Consortium. His research focuses o
n cohort epidemiology and implementation of multi-omic diagnostic tools fo
r personalized therapy of drug-resistant tuberculosis\, combining whole ge
nome sequencing\, expanded susceptibility testing\, and therapeutic drug m
onitoring to improve treatment outcomes while reducing side effects.\n \n▶
RECORDING [available here after event]\n\n \n Abstract \n\n
\n\n“Leveraging Modern Diagnostics to Personalize Treatment for Multidrug
Resistant Tuberculosis”\n\n \n \n▶RECORDING [available here after event]
DTSTART;TZID=America/New_York:20230502T103000
DTEND;TZID=America/New_York:20230502T113000
LOCATION:Gilman Hall: Gilman 50 (Marjorie Fisher Auditorium)
SEQUENCE:0
SUMMARY:“Leveraging Modern Diagnostics to Personalize Treatment for Multidr
ug Resistant Tuberculosis”
URL:https://icm.jhu.edu/events/leveraging-modern-diagnostics-to-personalize
-treatment-for-multidrug-resistant-tuberculosis/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2023/04/t
ornheim_headshot-scaled-e1682432235479.jpg\;259\;263\,medium\;https://icm.
jhu.edu/wp-content/uploads/2023/04/tornheim_headshot-scaled-e1682432235479
.jpg\;259\;263\,large\;https://icm.jhu.edu/wp-content/uploads/2023/04/torn
heim_headshot-scaled-e1682432235479.jpg\;259\;263\,full\;https://icm.jhu.e
du/wp-content/uploads/2023/04/tornheim_headshot-scaled-e1682432235479.jpg\
;259\;263
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Levera
ging Modern Diagnostics to Personalize Treatment for Multidrug Resistant T
uberculosis”
\n

\n
Jeff To
rnheim\, MD\, MPH is an Assistant Professor of Medicine\, Pediatrics\, and
International Health in the Division of Infectious Diseases at Johns Hopk
ins University and a TB clinician at the Baltimore City Health Department.
He completed a combined residency in Internal Medicine and Pediatrics at
the Yale University School of Medicine and a clinical fellowship in Infect
ious Diseases at the Johns Hopkins University School of Medicine before jo
ining the faculty in 2017. For the past 20 years he has engaged in clinica
l care\, physician education\, and translational research in India\, South
America\, and sub-Saharan Africa. He is a member of the AIDS Clinical Tri
als Group Tuberculosis Transformative Science Group\, the IMPAACT Network
TB Scientific Committee\, and a principal investigator in the RePORT India
Consortium. His research focuses on cohort epidemiology and implementatio
n of multi-omic diagnostic tools for personalized therapy of drug-resistan
t tuberculosis\, combining whole genome sequencing\, expanded susceptibili
ty testing\, and therapeutic drug monitoring to improve treatment outcomes
while reducing side effects.
\n
\n
▶RECORDING [available here after event]
\n
\n
\n Abstract
\n
\n
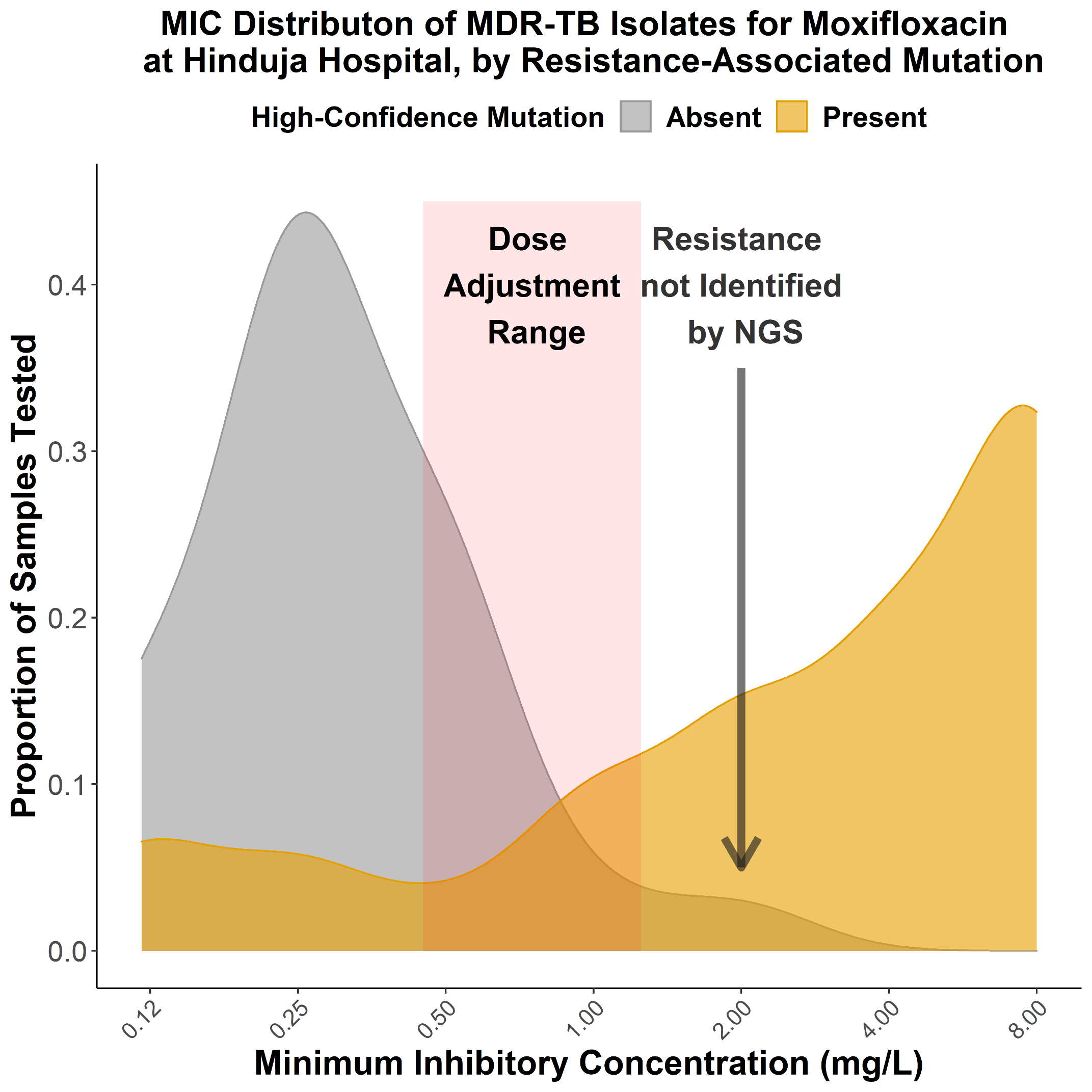

\n
\n
“Leveraging Modern Diagnostics to Pe
rsonalize Treatment for Multidrug Resistant Tuberculosis”
\n
\n
\n
\n
▶RECORDING [available here a
fter event]
\n
\n
\n
\n
\n
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984942@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series\,Events
CONTACT:Mishka Colombo\; 667-306-8941\; mcolomb4@jhu.edu\; https://wse.zoom
.us/j/97140116361
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Mathematical and Computational Frameworks for Adaptively Benchma
rking Patients in States of Health\, Disease\, and Recovery”\n\nDr. Brody
Foy is a research fellow in Systems Biology at Harvard Medical School and
Massachusetts General Hospital. He earned a BMath at Queensland University
of Technology\, and DPhil in Computer Science from the University of Oxfo
rd\, as a Rhodes Scholar. Dr Foy uses mathematical and computational appro
aches to quantify blood cell dynamics in acute and chronic disease setting
s. He is particularly interested in how we can better utilize routine clin
ical laboratory testing to generate physiologic and clinical insights. In
the winter he will be starting a lab at the University of Washington\, Dep
artment of Laboratory Medicine & Pathology\, as an acting Assistant Profes
sor.\n \n▶RECORDING [available here after event]\n\n \n Abstra
ct \n\n\n“Mathematical and Computational Frameworks for Adaptively Benc
hmarking Patients in States of Health\, Disease\, and Recovery”\nLaborator
y testing is a cornerstone of modern medicine. While cutting-edge assays a
re constantly in development\, the bulk of worldwide clinical testing is d
ominated by only a handful of markers. These ‘boring’ markers are regularl
y used in patient evaluation – but the physiologic insights they can provi
de are often overlooked. In this talk I will explore how mathematical and
statistical methods can be used to generate deep clinical and physiologic
insights from routine clinical laboratory tests such as the complete blood
count. From my own research I will show how careful analysis and modellin
g of biomarker dynamics can provide exciting and novel insights into homeo
static recovery and regulation\, chronic illness\, and physiologic shifts
such as pregnancy and menopause.\n \n▶RECORDING [available here after even
t]
DTSTART;TZID=America/New_York:20230905T160000
DTEND;TZID=America/New_York:20230905T171500
LOCATION:Clark 110
SEQUENCE:0
SUMMARY:Mathematical and Computational Frameworks for Adaptively Benchmarki
ng Patients in States of Health\, Disease\, and Recovery
URL:https://icm.jhu.edu/events/mathematical-and-computational-frameworks-fo
r-adaptively-benchmarking-patients-in-states-of-health-disease-and-recover
y/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2023/07/B
rody-Foy-Headshot-1.png\;219\;208\,medium\;https://icm.jhu.edu/wp-content/
uploads/2023/07/Brody-Foy-Headshot-1.png\;219\;208\,large\;https://icm.jhu
.edu/wp-content/uploads/2023/07/Brody-Foy-Headshot-1.png\;219\;208\,full\;
https://icm.jhu.edu/wp-content/uploads/2023/07/Brody-Foy-Headshot-1.png\;2
19\;208
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Mathem
atical and Computational Frameworks for Adaptively Benchmarking Patients i
n States of Health\, Disease\, and Recovery”<
/span>
\n

\n<
p>Dr. Brody Foy is a research fellow in Systems Biology at Harvard Medical
School and Massachusetts General Hospital. He earned a BMath at Queenslan
d University of Technology\, and DPhil in Computer Science from the Univer
sity of Oxford\, as a Rhodes Scholar. Dr Foy uses mathematical and computa
tional approaches to quantify blood cell dynamics in acute and chronic dis
ease settings. He is particularly interested in how we can better utilize
routine clinical laboratory testing to generate physiologic and clinical i
nsights. In the winter he will be starting a lab at the University of Wash
ington\, Department of Laboratory Medicine & Pathology\, as an acting Assi
stant Professor.\n
\n
▶RECORDI
NG [available here after event]
\n
\n
\n
Abstract
\n
\n


\n
“Mathematical and Comput
ational Frameworks for Adaptively Benchmarking Patients in States of Healt
h\, Disease\, and Recovery<
span style='font-size: 21pt\;'>”
\n
L
aboratory testing is a cornerstone of modern medicine. While cutting-edge
assays are constantly in development\, the bulk of worldwide clinical test
ing is dominated by only a handful of markers. These ‘boring’ markers are
regularly used in patient evaluation – but the physiologic insights they c
an provide are often overlooked. In this talk I will explore how mathemati
cal and statistical methods can be used to generate deep clinical and phys
iologic insights from routine clinical laboratory tests such as the comple
te blood count. From my own research I will show how careful analysis and
modelling of biomarker dynamics can provide exciting and novel insights in
to homeostatic recovery and regulation\, chronic illness\, and physiologic
shifts such as pregnancy and menopause.
\n
\n
▶RECORDING [available here after event]
\n\n
\n
\n \n
X-TAGS;LANGUAGE=en-US:Brody Foy\,Fall 2023\,ICM Distinguished Seminar Serie
s\,Seminar
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984963@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Events\,Tea Time
CONTACT:Mishka Colombo\; mcolomb4@jhu.edu
DESCRIPTION:ICM Tea Time\nPlease join us today for tea\, coffee\, snacks\,
and conversations with your fellow ICM colleagues.\nTuesdays 4:30PM-5:00PM
\nSecond floor hallway of Hackerman Hall\nClick Here to RSVP\n*Seminars wi
ll replace Team Time on the first Tuesday of each month during the academi
c calendar.
DTSTART;TZID=America/New_York:20230912T163000
DTEND;TZID=America/New_York:20230912T170000
EXRULE:FREQ=MONTHLY;BYDAY=1TU;WKST=MO
LOCATION:Hackerman Hall 2nd Floor Hallway
RRULE:FREQ=WEEKLY;BYDAY=TU;WKST=MO
SEQUENCE:0
SUMMARY:ICM Tea Time
URL:https://icm.jhu.edu/events/icm-tea-time/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2023/09/I
CM-Tea-Time-Sign.jpg\;781\;1078\,medium\;https://icm.jhu.edu/wp-content/up
loads/2023/09/ICM-Tea-Time-Sign.jpg\;781\;1078\,large\;https://icm.jhu.edu
/wp-content/uploads/2023/09/ICM-Tea-Time-Sign.jpg\;781\;1078\,full\;https:
//icm.jhu.edu/wp-content/uploads/2023/09/ICM-Tea-Time-Sign.jpg\;781\;1078
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
ICM <
/strong>Tea Time
\n<
p>Please join us today for tea\, coffee\, snacks\, and conversations with
your fellow ICM colleagues.\n
Tuesdays 4:30PM-5:00PM
\n
Second
floor hallway of Hackerman Hall
\n
Click Here to RSVP
\n
*Seminars will replace Team Time on t
he first Tuesday of each month during the academic calendar.![]()
\n
X-TAGS;LANGUAGE=en-US:Tea Time\,Tuesdays
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4984969@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series\,Special Seminars
CONTACT:Mishka Colombo\; 4105164116\; mcolomb4@jhu.edu\; https://wse.zoom.u
s/j/95676026583
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Leveraging Patient-Reported Outcome Dynamics to Predict Treatmen
t Response”\n\nDr. Renee Brady completed her postdoctoral training in the
Integrated Mathematical Oncology Department of the Houston Lee Moffitt Ca
ncer Center & Research Institute after earning her Bachelor of Science in
Mathematics from Florida A&M University and her Masters and PhD from North
Carolina State University. Her research focuses on developing novel\, pre
dictive models of non- and minimally-invasive biomarkers. After careful mo
del calibration and validation\, these models can be used to propose alter
native treatment strategies that can ultimately be used to reduce cancer h
ealth disparities.\n \n▶RECORDING\n\n \n Abstract \n\n\n“Le
veraging Patient-Reported Outcome Dynamics to Predict Treatment Response”
\nPatient-reported outcomes (PROs)\, collected using standardized question
naires at various time\npoints throughout a patient’s care\, provide an un
biased assessment of a patient’s health\ncondition\, reported directly by
the patient. Recent studies have shown that changes in PROs\nover time can
be early indicators of clinically important events such as cancer develop
ment and\nsurvival. While incredibly promising\, these studies fail to con
sider the patient-specific dynamics\nof individual PROs and how they might
be leveraged to predict individual patient responses to\ntreatment. This
is especially important in non-small cell lung cancer (NSCLC)\, which has
the\nlowest survival rates among all cancers. In this talk\, we demonstrat
e how PRO dynamics can be\nused as inter-radiographic predictors of tumor
volume changes. That is\, how PROs can be\nleveraged between radiographic
scans to predict tumor volume dynamics. This is assessed in\n108 NSCLC pat
ients receiving immune checkpoint inhibitors. The patients completed biwee
kly\nPRO questionnaires and received monthly tumor volume scans. We found
that changes in\nvolume were significantly correlated with dizziness (p
DTSTART;TZID=America/New_York:20231003T160000
DTEND;TZID=America/New_York:20231003T170000
LOCATION:Clark 110
SEQUENCE:0
SUMMARY:Leveraging Patient-Reported Outcome Dynamics to Predict Treatment
Response
URL:https://icm.jhu.edu/events/leveraging-patient-reported-outcome-dynamics
-to-predict-treatment-response/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2023/09/H
eadshot_Brady-2.jpg\;150\;188\,medium\;https://icm.jhu.edu/wp-content/uplo
ads/2023/09/Headshot_Brady-2.jpg\;150\;188\,large\;https://icm.jhu.edu/wp-
content/uploads/2023/09/Headshot_Brady-2.jpg\;150\;188\,full\;https://icm.
jhu.edu/wp-content/uploads/2023/09/Headshot_Brady-2.jpg\;150\;188
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Leveraging Patient-Reported Outcome
Dynamics to Predict Treatment Response”
\n

\n
Dr.
Renee Brady completed her postdoctoral training in the Integrated Mathem
atical Oncology Department of the Houston Lee Moffitt Cancer Center & Rese
arch Institute after earning her Bachelor of Science in Mathematics from F
lorida A&M University and her Masters and PhD from North Carolina State Un
iversity. Her research focuses on developing novel\, predictive models of
non- and minimally-invasive biomarkers. After careful model calibration an
d validation\, these models can be used to propose alternative treatment s
trategies that can ultimately be used to reduce cancer health disparities.
\n
\n
▶RECORDING
\n
\n
\n Abstract
\n
\n

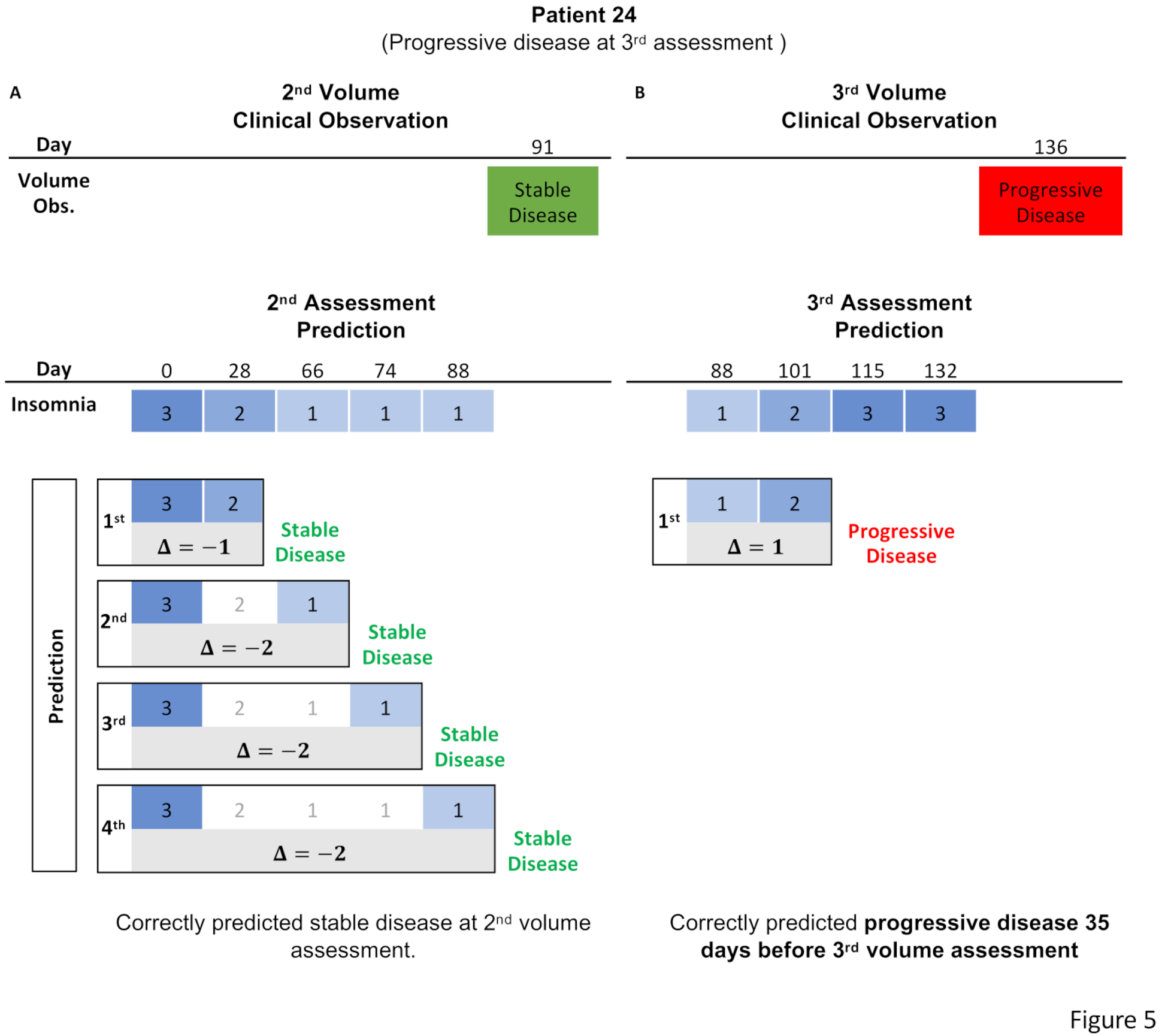
\n
“Leveraging P
atient-Reported Outcome Dynamics to Predict Treatment Response”
\n
Patient-reported outcomes (PROs)\, coll
ected using standardized questionnaires at various time
\npoints thro
ughout a patient’s care\, provide an unbiased assessment of a patient’s he
alth
\ncondition\, reported directly by the patient. Recent studies h
ave shown that changes in PROs
\nover time can be early indicators of
clinically important events such as cancer development and
\nsurviva
l. While incredibly promising\, these studies fail to consider the patient
-specific dynamics
\nof individual PROs and how they might be leverag
ed to predict individual patient responses to
\ntreatment. This is es
pecially important in non-small cell lung cancer (NSCLC)\, which has the\nlowest survival rates among all cancers. In this talk\, we demonstra
te how PRO dynamics can be
\nused as inter-radiographic predictors of
tumor volume changes. That is\, how PROs can be
\nleveraged between
radiographic scans to predict tumor volume dynamics. This is assessed in\n108 NSCLC patients receiving immune checkpoint inhibitors. The patie
nts completed biweekly
\nPRO questionnaires and received monthly tumo
r volume scans. We found that changes in
\nvolume were significantly
correlated with dizziness (p<0.005)\, insomnia (p <0.05)\, and fatigue
\n(p<0.05). Further analysis revealed that changes in insomnia could pre
dict progressive disease
\nwith a 77% accuracy\, with correct predict
ions of progressive disease occurring on average 45
\ndays prior to t
he next imaging study. Our study is an important first step in understandi
ng how
\nPROs can be utilized as a non-invasive and easily-obtained b
iomarker of when to change
\ntreatment to delay the development of tr
eatment progression.
\n
\n
▶RECORDING
\n\n
\n
\n \n
X-TAGS;LANGUAGE=en-US:Renee Brady\,Special Seminar
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4985033@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series\,Events
CONTACT:Mishka Colombo\; 4105164116\; mcolomb4@jhu.edu\; https://wse.zoom.u
s/j/98797164936
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Big Data Approaches to Study\nIntercellular Signaling during\nTu
mor Immune Evasion”\n\nDr. Peng Jiang started his research program at the
National Cancer Institute (NCI) in July 2019. His Lab focuses on developin
g big-data and artificial intelligence frameworks to identify biomarkers a
nd new therapeutic approaches for cancer immunotherapies in solid tumors.
Before joining NCI\, he finished his postdoctoral training at the Dana Far
ber Cancer Institute and Harvard University. During his postdoctoral resea
rch\, Peng developed computational frameworks that repurposed public domai
n data to identify biomarkers and regulators of cancer immunotherapy resis
tance. Notably\, his computational model TIDE revealed that cancer cells c
ould utilize the self-protection strategy of cytotoxic lymphocytes to resi
st lymphocyte killing under immune checkpoint blockade. Dr. Peng finished
his Ph.D. at the Department of Computer Science & Lewis Sigler Genomics In
stitute at Princeton University\, and his undergraduate study with the hig
hest national honors at the Department of Computer Science at Tsinghua Uni
versity (GPA rank 1st in his year). He is a recipient of the NCI K99 Pathw
ay to Independence Award\, the Scholar-In-Training Award of the American A
ssociation of Cancer Research\, and the Technology Innovation Award of the
Cancer Research Institute.\n \n▶RECORDING \n\n \n Abstract
\n\n\n“Big Data Approaches to Study\nIntercellular Signaling during\nTumo
r Immune Evasion”\nMy talk will present three computational frameworks we
developed to study cytokine signaling activities and cell-cell communicati
ons during the antitumor immune response. The basic immunology tool to stu
dy cytokine signaling mostly measures cytokine release\, which is transien
t and does not represent downstream target activities. Therefore\, we firs
t developed the CytoSig platform\, providing a database of target genes mo
dulated by cytokines and a predictive model of cytokine signaling cascades
from transcriptomic profiles. We collected 20\,591 transcriptome profiles
for human cytokine\, chemokine\, and growth factor responses. This atlas
of transcriptional patterns induced by cytokines enabled the reliable pred
iction of signaling activities in distinct cell populations in infectious
diseases\, chronic inflammation\, and cancer using bulk and single-cell tr
anscriptomic data. CytoSig revealed previously unidentified roles of many
cytokines\, such as BMP6 as an anti-inflammatory factor. Then\, based on C
ytoSig\, we developed Tres\, a computational model utilizing single-cell t
ranscriptomic data to identify signatures of T cells that are resilient to
immunosuppressive signals. Tres reliably predicts clinical responses to i
mmunotherapy in multiple cancer types using bulk T cell transcriptomic dat
a from pre-treatment patient tumors or infusion/pre-manufacture samples fo
r cellular immunotherapies. Further\, Tres identified FIBP as a candidate
immunotherapy target to potentiate adoptive cell therapy efficacies. Last\
, I will briefly show our SpaCET framework for deconvolving cell fractions
and identifying cell-cell interactions in tumor spatial transcriptomics d
ata.\n \n▶RECORDING
DTSTART;TZID=America/New_York:20231107T160000
DTEND;TZID=America/New_York:20231107T170000
LOCATION:Clark 110
SEQUENCE:0
SUMMARY:Big Data Approaches to Study Intercellular Signaling during Tumor I
mmune Evasion
URL:https://icm.jhu.edu/events/big-data-approaches-to-study-intercellular-s
ignaling-during-tumor-immune-evasion/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2023/10/1
1-7-23-Peng-Jiang-Headshot-1.jpg\;232\;232\,medium\;https://icm.jhu.edu/wp
-content/uploads/2023/10/11-7-23-Peng-Jiang-Headshot-1.jpg\;232\;232\,larg
e\;https://icm.jhu.edu/wp-content/uploads/2023/10/11-7-23-Peng-Jiang-Heads
hot-1.jpg\;232\;232\,full\;https://icm.jhu.edu/wp-content/uploads/2023/10/
11-7-23-Peng-Jiang-Headshot-1.jpg\;232\;232
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Big Da
ta Approaches to Study
\nIntercellular Signaling during
\nTumor
Immune Evasion”
\n

\n
Dr. Peng Jiang started his rese
arch program at the National Cancer Institute (NCI) in July 2019. His Lab
focuses on developing big-data and artificial intelligence frameworks to i
dentify biomarkers and new therapeutic approaches for cancer immunotherapi
es in solid tumors. Before joining NCI\, he finished his postdoctoral trai
ning at the Dana Farber Cancer Institute and Harvard University. During hi
s postdoctoral research\, Peng developed computational frameworks that rep
urposed public domain data to identify biomarkers and regulators of cancer
immunotherapy resistance. Notably\, his computational model TIDE revealed
that cancer cells could utilize the self-protection strategy of cytotoxic
lymphocytes to resist lymphocyte killing under immune checkpoint blockade
. Dr. Peng finished his Ph.D. at the Department of Computer Science & Lewi
s Sigler Genomics Institute at Princeton University\, and his undergraduat
e study with the highest national honors at the Department of Computer Sci
ence at Tsinghua University (GPA rank 1st in his year). He is a recipient
of the NCI K99 Pathway to Independence Award\, the Scholar-In-Training Awa
rd of the American Association of Cancer Research\, and the Technology Inn
ovation Award of the Cancer Research Institute.
\n
\n
▶RECORDING
\n
\n
\n Abstract
\n
\n

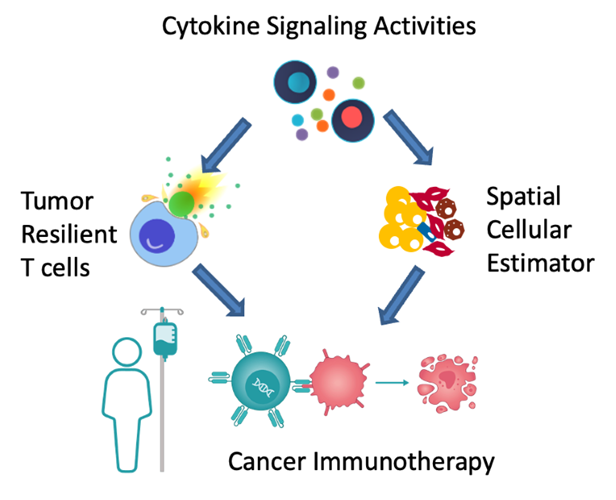
\n
“B
ig Data Approaches to Study
\nIntercellular Signaling during
\nT
umor Immune Evasion”
\n
My talk
will present three computational frameworks we developed to study cytokine
signaling activities and cell-cell communications during the antitumor im
mune response. The basic immunology tool to study cytokine signaling mostl
y measures cytokine release\, which is transient and does not represent do
wnstream target activities. Therefore\, we first developed the CytoSig pla
tform\, providing a database of target genes modulated by cytokines and a
predictive model of cytokine signaling cascades from transcriptomic profil
es. We collected 20\,591 transcriptome profiles for human cytokine\, chemo
kine\, and growth factor responses. This atlas of transcriptional patterns
induced by cytokines enabled the reliable prediction of signaling activit
ies in distinct cell populations in infectious diseases\, chronic inflamma
tion\, and cancer using bulk and single-cell transcriptomic data. CytoSig
revealed previously unidentified roles of many cytokines\, such as BMP6 as
an anti-inflammatory factor. Then\, based on CytoSig\, we developed Tres\
, a computational model utilizing single-cell transcriptomic data to ident
ify signatures of T cells that are resilient to immunosuppressive signals.
Tres reliably predicts clinical responses to immunotherapy in multiple ca
ncer types using bulk T cell transcriptomic data from pre-treatment patien
t tumors or infusion/pre-manufacture samples for cellular immunotherapies.
Further\, Tres identified FIBP as a candidate immunotherapy target to pot
entiate adoptive cell therapy efficacies. Last\, I will briefly show our S
paCET framework for deconvolving cell fractions and identifying cell-cell
interactions in tumor spatial transcriptomics data.
\n
\n
▶RECORDING
\n
\n
\n
\n \n
X-TAGS;LANGUAGE=en-US:Distinguished Seminar Series\,Peng Jiang
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4985055@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series\,Events
CONTACT:Mishka Colombo\; 667-306-8941\; mcolomb4@jhu.edu\; https://wse.zoom
.us/j/94073678803
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Predicting Immunogenic\nNeoepitopes in Cancer”\n\nDr. Rachel Kar
chin is a Professor in the Departments of Biomedical Engineering and Oncol
ogy\, with a secondary appointment in Computer Science at Johns Hopkins Un
iversity\, She received a Ph.D. in Computer Science from the University of
California\, Santa Cruz in 2003\, spent three years as a postdoctoral fel
low in the Department of Biopharmaceutical Sciences at University of Calif
ornia\, San Francisco\, and joined the Hopkins faculty in 2006. Working cl
osely with cancer geneticists\, pathologists and oncologists\, her lab has
developed novel tools to identify pathogenic missense mutations and drive
r genes\, to model tumor evolution from next-generation sequencing data\,
and to predict tumor neoepitopes.\n \n▶RECORDING \n\n \n Abst
ract \n\n\n“Predicting Immunogenic\nNeoepitopes in Cancer”\nIdentifying
neoepitopes that elicit an adaptive immune response is a major bottleneck
to developing personalized cancer vaccines. Experimental validation of ca
ndidate neoepitopes is extremely resource intensive and the vast majority
of candidates are non-immunogenic\, creating a needle-in-a-haystack proble
m. Here we address this challenge\, presenting computational methods for p
redicting class I major histocompatibility complex (MHC-I) epitopes and id
entifying immunogenic neoepitopes with improved precision. The BigMHC meth
od comprises an ensemble of seven pan-allelic deep neural networks trained
on peptide-MHC eluted ligand data from mass spectrometry assays and trans
fer learned on data from assays of antigen-specific immune response. Compa
red with four state-of-the-art classifiers\, BigMHC significantly improves
the prediction of epitope presentation on a test set of 45\,409 MHC ligan
ds among 900\,592 random negatives (area under the receiver operating char
acteristic = 0.9733\; area under the precision-recall curve = 0.8779). Aft
er transfer learning on immunogenicity data\, BigMHC yields significantly
higher precision than seven state-of-the-art models in identifying immunog
enic neoepitopes\, making BigMHC effective in clinical settings.\n \n▶RECO
RDING
DTSTART;TZID=America/New_York:20231205T160000
DTEND;TZID=America/New_York:20231205T170000
LOCATION:Clark 110
SEQUENCE:0
SUMMARY:Predicting Immunogenic Neoepitopes in Cancer
URL:https://icm.jhu.edu/events/predicting-immunogenic-neoepitopes-in-cancer
/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2023/11/R
achel-Headshot-Photo.jpg\;300\;300\,medium\;https://icm.jhu.edu/wp-content
/uploads/2023/11/Rachel-Headshot-Photo.jpg\;300\;300\,large\;https://icm.j
hu.edu/wp-content/uploads/2023/11/Rachel-Headshot-Photo.jpg\;300\;300\,ful
l\;https://icm.jhu.edu/wp-content/uploads/2023/11/Rachel-Headshot-Photo.jp
g\;300\;300
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Predic
ting Immunogenic
\nNeoepitopes in Cancer”
\n

\n
Dr. Rachel Karchin is a Professor in the Departments
of Biomedical Engineering and Oncology\, with a secondary appointment in C
omputer Science at Johns Hopkins University\, She received a Ph.D. in Comp
uter Science from the University of California\, Santa Cruz in 2003\, spen
t three years as a postdoctoral fellow in the Department of Biopharmaceuti
cal Sciences at University of California\, San Francisco\, and joined the
Hopkins faculty in 2006. Working closely with cancer geneticists\, patholo
gists and oncologists\, her lab has developed novel tools to identify path
ogenic missense mutations and driver genes\, to model tumor evolution from
next-generation sequencing data\, and to predict tumor neoepitopes.
\n
\n
▶RECORDING
\n
\n
\n Abstract
\n
\n

 \n
\n“Predicting Immunogenic
\nNeoepitopes in Cancer”
\n
Identifying neoepitopes that
elicit an adaptive immune response is a major bottleneck to developing per
sonalized cancer vaccines. Experimental validation of candidate neoepitope
s is extremely resource intensive and the vast majority of candidates are
non-immunogenic\, creating a needle-in-a-haystack problem. Here we address
this challenge\, presenting computational methods for predicting class I
major histocompatibility complex (MHC-I) epitopes and identifying immunoge
nic neoepitopes with improved precision. The BigMHC method comprises an en
semble of seven pan-allelic deep neural networks trained on peptide-MHC el
uted ligand data from mass spectrometry assays and transfer learned on dat
a from assays of antigen-specific immune response. Compared with four stat
e-of-the-art classifiers\, BigMHC significantly improves the prediction of
epitope presentation on a test set of 45\,409 MHC ligands among 900\,592
random negatives (area under the receiver operating characteristic = 0.973
3\; area under the precision-recall curve = 0.8779). After transfer learni
ng on immunogenicity data\, BigMHC yields significantly higher precision t
han seven state-of-the-art models in identifying immunogenic neoepitopes\,
making BigMHC effective in clinical settings.
\n
\n
▶RECORDING
\n
\n
\n \n \n
X-TAGS;LANGUAGE=en-US:Rachel Karchin\,Seminar
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4985083@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series
CONTACT:Mishka Colombo\; 4105164116\; mcolomb4@jhu.edu\; https://wse.zoom.u
s/j/99695749134
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Mathematical and Computational Frameworks for Adaptively Benchma
rking Patients in States of Health\, Disease\, and Recovery”\n\nDr. Brody
Foy is a junior faculty member at the University of Washington\, Departmen
t of Laboratory Medicine & Pathology. Brody completed his DPhil in Compute
r Science at the University of Oxford\, as a Rhodes Scholar\, and undertoo
k postdoctoral training at Harvard Medical School & Massachusetts General
Hospital. A mathematician by training\, Brody’s research is focused on usi
ng math modelling and machine learning to improve utilization of clinical
laboratory data\, and increase the quality of information extraction from
blood testing. His lab uses computational tools to learn about human physi
ology\, improve clinical workflows\, and develop novel tools for patient c
are.\n \n▶RECORDING \n\n \n Abstract \n\n\n“Mathematical an
d Computational Frameworks for Adaptively Benchmarking Patients in States
of Health\, Disease\, and Recovery”\nLaboratory testing is a cornerstone o
f modern medicine. While cutting-edge assays are constantly in development
\, the bulk of worldwide clinical testing is dominated by only a handful o
f markers. These ‘boring’ markers are regularly used in patient evaluation
– but the physiologic insights they can provide are often overlooked. In
this talk I will explore how mathematical and statistical methods can be u
sed to generate deep clinical and physiologic insights from routine clinic
al laboratory tests such as the complete blood count. From my own research
I will show how careful analysis and modelling of biomarker dynamics can
provide exciting and novel insights into homeostatic recovery and regulati
on\, chronic illness\, and physiologic shifts such as pregnancy and menopa
use.\n \n▶RECORDING
DTSTART;TZID=America/New_York:20240305T160000
DTEND;TZID=America/New_York:20240305T170000
LOCATION:Clark 110
SEQUENCE:0
SUMMARY:Mathematical and Computational Frameworks for Adaptively Benchmarki
ng Patients in States of Health\, Disease\, and Recovery
URL:https://icm.jhu.edu/events/mathematical-and-computational-frameworks-fo
r-adaptively-benchmarking-patients-in-states-of-health-disease-and-recover
y-2/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2024/01/B
rody-Foy_Cropped.jpg\;220\;220\,medium\;https://icm.jhu.edu/wp-content/upl
oads/2024/01/Brody-Foy_Cropped.jpg\;220\;220\,large\;https://icm.jhu.edu/w
p-content/uploads/2024/01/Brody-Foy_Cropped.jpg\;220\;220\,full\;https://i
cm.jhu.edu/wp-content/uploads/2024/01/Brody-Foy_Cropped.jpg\;220\;220
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Mathem
atical and Computational Frameworks for Adaptively Benchmarking Patients i
n States of Health\, Disease\, and Recovery”<
/span>
\n

\n
D
r. Brody Foy is a junior faculty member at the University of Washington\,
Department of Laboratory Medicine & Pathology. Brody completed his DPhil i
n Computer Science at the University of Oxford\, as a Rhodes Scholar\, and
undertook postdoctoral training at Harvard Medical School & Massachusetts
General Hospital. A mathematician by training\, Brody’s research is focus
ed on using math modelling and machine learning to improve utilization of
clinical laboratory data\, and increase the quality of information extract
ion from blood testing. His lab uses computational tools to learn about hu
man physiology\, improve clinical workflows\, and develop novel tools for
patient care.
\n
\n
▶RECORDING
\n
\n
\n Abstract
\n
\n


\n
<
strong>“Mathematical and Computational Frameworks for Adaptively Benchmark
ing Patients in States of Health\, Disease\, and Recovery<
span style='color: #000000\;'>”
\n
Laboratory testing is a cornerstone of moder
n medicine. While cutting-edge assays are constantly in development\, the
bulk of worldwide clinical testing is dominated by only a handful of marke
rs. These ‘boring’ markers are regularly used in patient evaluation – but
the physiologic insights they can provide are often overlooked. In this ta
lk I will explore how mathematical and statistical methods can be used to
generate deep clinical and physiologic insights from routine clinical labo
ratory tests such as the complete blood count. From my own research I will
show how careful analysis and modelling of biomarker dynamics can provide
exciting and novel insights into homeostatic recovery and regulation\, ch
ronic illness\, and physiologic shifts such as pregnancy and menopause.
\n
\n
▶RECORDING
\n
\n
\n
\n \n
X-TAGS;LANGUAGE=en-US:Brody Foy
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4985143@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES;LANGUAGE=en-US:Distinguished Seminar Series\,Events
CONTACT:Mishka Colombo\; 667-306-8941\; mcolomb4@jhu.edu\; https://wse.zoom
.us/j/97571700790
DESCRIPTION:Jump to:\n \n \n Bio\n \n
\n Abstract\n \n \n \n \n
Bio \n“Overcoming Analytic Challenges in Microbiome Science”\nUMIACS -D
r. Mihai Pop\nMihai Pop is a professor of computer science and director of
the University of Maryland\nInstitute for Advanced Computer Studies. He d
evelops computational approaches for analyzing\nmicrobial communities\, pa
rticularly for characterizing their strain-level diversity. Other\ninteres
ts include biological databases\, antibiotic resistance\, and software tes
ting. His lab\nhas developed several widely used open-source software tool
s for the analysis of genomic and\nmetagenomic data. Pop teaches at all ac
ademic professional levels\, and is particularly\ninterested in developing
open educational resources for introductory computer science and\nbioinfo
rmatics. He strongly advocates for inclusion and diversity within the scie
ntific\ncommunity. Pop completed his undergraduate studies in 1994 at the
Politehnica University in\nBucharest\, Romania\, received his Ph.D. in com
puter science from Johns Hopkins University in 2000\,\nand has been at the
University of Maryland since 2005. He is a fellow of the Association of\n
Computing Machinery and of the International Society for Computational Bio
logy.\n \n▶RECORDING\n\n \n Abstract \n\n\n“Overcoming Anal
ytic Challenges in Microbiome Science”\n \nAs microbiome research matures\
, it has become clear that a better understanding of the\nmicrobial commun
ities inhabiting our world is key to a better understanding of our\nenviro
nment and of animal and human health. At the same time\, we have become aw
are of\nthe limitations current microbiome technologies have\, and of the
tremendous challenges\nposed by the analysis of the massive data sets gene
rated in microbiome studies. In my\ntalk I will describe some of the resea
rch taking place in my lab aimed at developing\ncomputational tools for mi
crobiome analyses. I will specifically focus on challenges\nrelated to the
structure of biological databases\, and the resulting impact on the\ninsi
ghts that can be derived from microbiome data.\n \n▶RECORDING
DTSTART;TZID=America/New_York:20240402T160000
DTEND;TZID=America/New_York:20240402T170000
SEQUENCE:0
SUMMARY:Overcoming Analytic Challenges in Microbiome Science
URL:https://icm.jhu.edu/events/overcoming-analytic-challenges-in-microbiome
-science/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2024/01/M
ihai-Pop.jpg\;220\;220\,medium\;https://icm.jhu.edu/wp-content/uploads/202
4/01/Mihai-Pop.jpg\;220\;220\,large\;https://icm.jhu.edu/wp-content/upload
s/2024/01/Mihai-Pop.jpg\;220\;220\,full\;https://icm.jhu.edu/wp-content/up
loads/2024/01/Mihai-Pop.jpg\;220\;220
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
\n
Jump to:
\n
\n
\n
\n Bio
\n
“Overco
ming Analytic Challenges in Microbiome Science”
\n

UMIACS -Dr. Mihai P
op
\n
Mihai Pop is a professor
of computer science and director of the University of Maryland
\nIns
titute for Advanced Computer Studies. He develops computational approaches
for analyzing
\nmicrobial communities\, particularly for characteriz
ing their strain-level diversity. Other
\ninterests include biologica
l databases\, antibiotic resistance\, and software testing. His lab
\nhas developed several widely used open-source software tools for the ana
lysis of genomic and
\nmetagenomic data. Pop teaches at all academic
professional levels\, and is particularly
\ninterested in developing
open educational resources for introductory computer science and
\nbi
oinformatics. He strongly advocates for inclusion and diversity within the
scientific
\ncommunity. Pop completed his undergraduate studies in 1
994 at the Politehnica University in
\nBucharest\, Romania\, received
his Ph.D. in computer science from Johns Hopkins University in 2000\,
\nand has been at the University of Maryland since 2005. He is a fellow
of the Association of
\nComputing Machinery and of the International
Society for Computational Biology.
\n
\n
▶RECORDING
\n
\n
\n Abstract
\n
\n

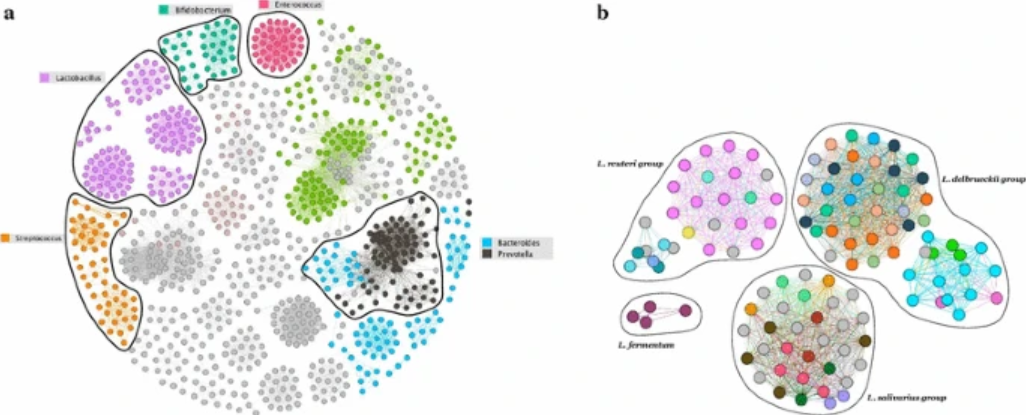 \n
\n“Overcoming Analytic Challenges in Microbiome Science
”
\n
\n
As microbiome research matures\, it has become clear that a be
tter understanding of the
\nmicrobial communities inhabiting our worl
d is key to a better understanding of our
\nenvironment and of animal
and human health. At the same time\, we have become aware of
\nthe l
imitations current microbiome technologies have\, and of the tremendous ch
allenges
\nposed by the analysis of the massive data sets generated i
n microbiome studies. In my
\ntalk I will describe some of the resear
ch taking place in my lab aimed at developing
\ncomputational tools f
or microbiome analyses. I will specifically focus on challenges
\nrel
ated to the structure of biological databases\, and the resulting impact o
n the
\ninsights that can be derived from microbiome data.
\n
<
/p>\n
▶RECOR
DING
\n
\n
\n
\n \n
X-TAGS;LANGUAGE=en-US:Mihai Pop
END:VEVENT
BEGIN:VEVENT
UID:ai1ec-4985119@icm.jhu.edu
DTSTAMP:20240727T143135Z
CATEGORIES:
CONTACT:Sabrina Sengupta\; 410-516-6892\; ssengu19@jhu.edu\; https://forms.
gle/jnG6nmcSgkZNMWmp8
DESCRIPTION:Register for the event by clicking here.
DTSTART;TZID=America/New_York:20240402T170000
DTEND;TZID=America/New_York:20240402T193000
LOCATION:Hackerman B17
SEQUENCE:0
SUMMARY:CM Night
URL:https://icm.jhu.edu/events/cm-night-2/
X-COST-TYPE:free
X-WP-IMAGES-URL:thumbnail\;https://icm.jhu.edu/wp-content/uploads/2024/03/C
M-Night-Flyer-2024-002-1-791x1024.png\;791\;1024\,medium\;https://icm.jhu.
edu/wp-content/uploads/2024/03/CM-Night-Flyer-2024-002-1-791x1024.png\;791
\;1024\,large\;https://icm.jhu.edu/wp-content/uploads/2024/03/CM-Night-Fly
er-2024-002-1-791x1024.png\;791\;1024\,full\;https://icm.jhu.edu/wp-conten
t/uploads/2024/03/CM-Night-Flyer-2024-002-1-791x1024.png\;791\;1024
X-ALT-DESC;FMTTYPE=text/html:\\n\\n\\n
\\n\\n
Register for
the event by clicking here. 
\n
END:VEVENT
END:VCALENDAR