Bio
“Allele-specific Stochasticity of the Epigenome”

Aleksandar Milosavljevic, PhD., is a Henry and Emma Meyer Professor of Molecular and Human Genetics and Director of the Program in Quantitative and Computational Biosciences at Baylor College of Medicine. Research in Dr. Milosavljevic’s Bioinformatics Research Laboratory (BRL) includes bioinformatics, genomics, clinical genomics, epigenomics, cancer biology and extracellular RNA communication
To join live event click here: https://wse.zoom.us/j/96447421269
 Recording will be available here after the event.
Recording will be available here after the event.
Abstract
“Allele-specific Stochasticity of the Epigenome”
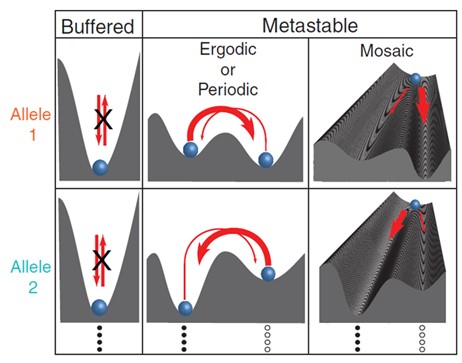
Stochasticity is a long-recognized characteristic of the epigenome, tracing back to the discovery of random X-chromosome inactivation and random allelic expression of olfactory and immunity-related genes. Allele-specific DNA methylation, transcription and chromatin marks have long been recognized as hallmarks of imprinted loci. More recently, ubiquitous sequence-dependent allele-specific DNA methylation and chromatin marks came to light. Our most recent analyses based on high-resolution allele-specific epigenomic maps from the NIH Roadmap Epigenome and ENTEx projects reveal sequence-dependent stochastic switching at gene regulatory loci mediated by transcription-factor binding. The analyses provide a unifying model that links sequence-dependent allelic imbalances of the epigenome, stochastic switching at gene regulatory loci, selective buffering of the regulatory circuitry against the effects of random mutations, and disease-associated genetic variation. We review these findings and explore their relevance for interpreting the genetic variation in regulatory regions revealed by whole-genome sequencing.
 Recording will be available here after the event.
Recording will be available here after the event.