Computational Physiological Medicine
The goal of computational physiological medicine is to develop mechanistic models of biological systems in disease, personalize these models using patient data, and apply them to improve disease diagnosis and treatment.
Understanding disease and its treatment requires development of models spanning multiple levels of biological organization. Computational physiological medicine seeks to develop models of disease that integrate information from the level of molecular networks to cells, tissues, organs and organ systems. This multi-scale modeling approach has applications to cancer, diabetes, heart and brain disease, and others. Computational models are typically developed using data obtained from animal experimental models of disease, and are then specialized using more limited human data sets. To the greatest extent possible, experiments and modeling proceed in a highly modular fashion in which distinct biological processes are teased apart and characterized. Models components are then brought together and tested to see if they are able to reproduce and predict novel experimental data.
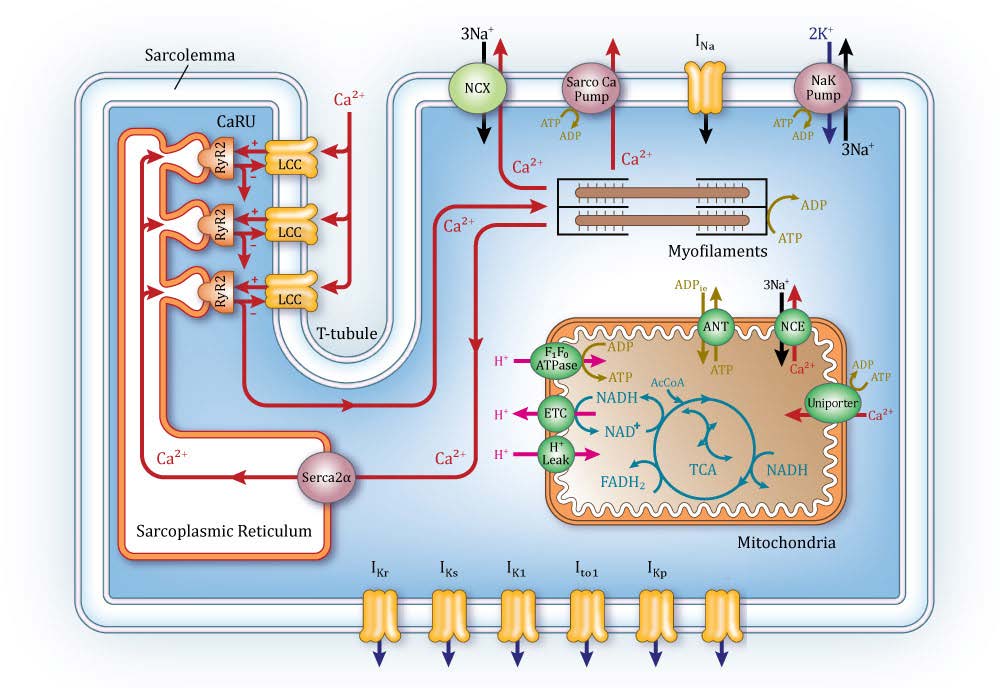
Among computational models of various physiological systems, the heart is one of the most highly advanced examples of a “virtual organ.” There is a long history of cardiac modeling, beginning more than 50 years ago with publication of the first model of the cardiac myocyte action potential. Since then, myocyte modeling has progressed rapidly by incorporating descriptions of many different subcellular processes and the ways they regulate properties such as the action potential. Cardiac modeling has also progressed to the level of the whole heart. Whole-heart image-based models take the form of reaction-diffusion partial differential equations, where the reaction term is specified by the system of equations modeling the myocyte, and the diffusion term is specified by the image-based measurements of heart anatomy along with estimated cell-to-cell coupling by gap junctions. This has led to a new generation of whole-heart image-based models with unprecedented structural and biophysical detail, including cardiac electromechanics, and even models integrating from the levels of cellular electrophysiology to electromechanics, to fluid dynamics of blood flow within the left ventricle. ICM researchers apply similar approaches to study brain disease, HIV and other immune system disorders, and personalized pharmacotherapy.